Isolation of a Potently Neutralizing and Protective Human Monoclonal Antibody Targeting Yellow Fever Virus
- PMID: 35420472
- PMCID: PMC9239089
- DOI: 10.1128/mbio.00512-22
Isolation of a Potently Neutralizing and Protective Human Monoclonal Antibody Targeting Yellow Fever Virus
Abstract
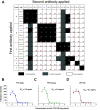
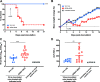
Yellow fever virus (YFV) causes sporadic outbreaks of infection in South America and sub-Saharan Africa. While live-attenuated yellow fever virus vaccines based on three substrains of 17D are considered some of the most effective vaccines in use, problems with production and distribution have created large populations of unvaccinated, vulnerable individuals in areas of endemicity. To date, specific antiviral therapeutics have not been licensed for human use against YFV or any other related flavivirus. Recent advances in monoclonal antibody (mAb) technology have allowed the identification of numerous candidate therapeutics targeting highly pathogenic viruses, including many flaviviruses. Here, we sought to identify a highly neutralizing antibody targeting the YFV envelope (E) protein as a therapeutic candidate. We used human B cell hybridoma technology to isolate mAbs from circulating memory B cells from human YFV vaccine recipients. These antibodies bound to recombinant YFV E protein and recognized at least five major antigenic sites on E. Two mAbs (designated YFV-136 and YFV-121) recognized a shared antigenic site and neutralized the YFV-17D vaccine strain in vitro. YFV-136 also potently inhibited infection by multiple wild-type YFV strains, in part, at a postattachment step in the virus replication cycle. YFV-136 showed therapeutic protection in two animal models of YFV challenge, including hamsters and immunocompromised mice engrafted with human hepatocytes. These studies define features of the antigenic landscape of the YFV E protein recognized by the human B cell response and identify a therapeutic antibody candidate that inhibits infection and disease caused by highly virulent strains of YFV. IMPORTANCE Yellow fever virus (YFV) is a mosquito-borne virus that occasionally causes outbreaks of severe infection and disease in South America and sub-Saharan Africa. There are very effective live-attenuated (weakened) yellow fever virus vaccines, but recent problems with their production and distribution have left many people in affected areas vulnerable. Here, we sought to isolate an antibody targeting the surface of the virus for possible use in the future as a biologic drug to prevent or treat YFV infection. We isolated naturally occurring antibodies from individuals who had received a YFV vaccine. We created antibodies and tested them. We found that the antibody with the most powerful antiviral activity was a beneficial treatment in two different small-animal models of human infection. These studies identified features of the virus that are recognized by the human immune system and generated a therapeutic antibody candidate that inhibits infection caused by highly virulent strains of YFV.
Keywords: monoclonal antibodies; mouse model; neutralization; vaccine; yellow fever virus.
Conflict of interest statement
The authors declare a conflict of interest. M.L.G. is an unpaid member of the scientific advisory boards of Protein Metrics and GenNext. M.S.D. is a consultant for Inbios, Vir Biotechnology, Senda Biosciences, and Carnival Corporation and on the scientific advisory boards of Moderna and Immunome. The Diamond laboratory has received funding support in sponsored research agreements from Moderna, Vir Biotechnology, and Emergent BioSolutions. J.E.C. has served as a consultant for Luna Innovations, Merck, GlaxoSmithKline, is a member of the scientific advisory board of Meissa Vaccines and is Founder of IDBiologics. The Crowe laboratory at Vanderbilt University Medical Center has received unrelated sponsored research agreements from IDBiologics, AstraZeneca and Takeda.
Figures




Comment in
-
Monoclonal Antibodies and Flaviviruses: a Possible Option?mBio. 2022 Jun 28;13(3):e0082422. doi: 10.1128/mbio.00824-22. Epub 2022 May 16. mBio. 2022. PMID: 35575500 Free PMC article.
References
-
- Cunha MS, Tubaki RM, de Menezes RMT, Pereira M, Caleiro GS, Coelho E, Saad LDC, Fernandes NCCDA, Guerra JM, Nogueira JS, Summa JL, Coimbra AAC, Zwarg T, Witkin SS, Mucci LF, Timenetsky MDCST, Sabino EC, de Deus JT. 2020. Possible non-sylvatic transmission of yellow fever between non-human primates in Sao Paulo city, Brazil, 2017-2018. Sci Rep 10:15751. doi:10.1038/s41598-020-72794-x. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

