The DP5 probability, quantification and visualisation of structural uncertainty in single molecules
- PMID: 35432857
- PMCID: PMC8943899
- DOI: 10.1039/d1sc04406k
The DP5 probability, quantification and visualisation of structural uncertainty in single molecules
Abstract
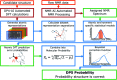
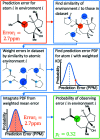
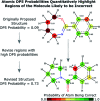
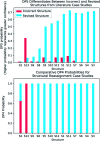
Whenever a new molecule is made, a chemist will justify the proposed structure by analysing the NMR spectra. The widely-used DP4 algorithm will choose the best match from a series of possibilities, but draws no conclusions from a single candidate structure. Here we present the DP5 probability, a step-change in the quantification of molecular uncertainty: given one structure and one 13C NMR spectra, DP5 gives the probability of the structure being correct. We show the DP5 probability can rapidly differentiate between structure proposals indistinguishable by NMR to an expert chemist. We also show in a number of challenging examples the DP5 probability may prevent incorrect structures being published and later reassigned. DP5 will prove extremely valuable in fields such as discovery-driven automated chemical synthesis and drug development. Alongside the DP4-AI package, DP5 can help guide synthetic chemists when resolving the most subtle structural uncertainty. The DP5 system is available at https://github.com/Goodman-lab/DP5.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures


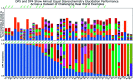
References
-
- Kutateladze A. G. Holt T. J. Org. Chem. 2019;84:51. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

