Discovery of Potent Orally Bioavailable WD Repeat Domain 5 (WDR5) Inhibitors Using a Pharmacophore-Based Optimization
- PMID: 35436124
- PMCID: PMC10081510
- DOI: 10.1021/acs.jmedchem.2c00195
Discovery of Potent Orally Bioavailable WD Repeat Domain 5 (WDR5) Inhibitors Using a Pharmacophore-Based Optimization
Abstract
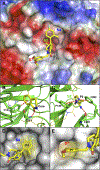
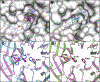
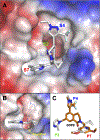
WD repeat domain 5 (WDR5) is a nuclear scaffolding protein that forms many biologically important multiprotein complexes. The WIN site of WDR5 represents a promising pharmacological target in a variety of human cancers. Here, we describe the optimization of our initial WDR5 WIN-site inhibitor using a structure-guided pharmacophore-based convergent strategy to improve its druglike properties and pharmacokinetic profile. The core of the previous lead remained constant while a focused SAR effort on the three pharmacophore units was combined to generate a new in vivo lead series. Importantly, this new series of compounds has picomolar binding affinity, improved cellular antiproliferative activity and selectivity, and increased kinetic aqueous solubility. They also exhibit a desirable oral pharmacokinetic profile with manageable intravenous clearance and high oral bioavailability. Thus, these new leads are useful probes toward studying the effects of WDR5 inhibition.
Conflict of interest statement
The authors declare the following competing financial interest(s): K.B.T., K.M.M., J.J.M., J.T., J.A., T.L., and S.W.F. are inventors on a patent application that was filed to protect this series of WDR5 inhibitors. All other authors declare no competing interest.
Figures
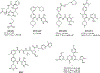


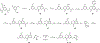
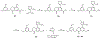
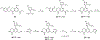
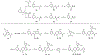
Similar articles
-
Structure-based discovery of potent WD repeat domain 5 inhibitors that demonstrate efficacy and safety in preclinical animal models.Proc Natl Acad Sci U S A. 2023 Jan 3;120(1):e2211297120. doi: 10.1073/pnas.2211297120. Epub 2022 Dec 27. Proc Natl Acad Sci U S A. 2023. PMID: 36574664 Free PMC article.
-
Discovery and Structure-Based Optimization of Potent and Selective WD Repeat Domain 5 (WDR5) Inhibitors Containing a Dihydroisoquinolinone Bicyclic Core.J Med Chem. 2020 Jan 23;63(2):656-675. doi: 10.1021/acs.jmedchem.9b01608. Epub 2020 Jan 7. J Med Chem. 2020. PMID: 31858797 Free PMC article.
-
Structure-Based Discovery of Potent, Orally Bioavailable Benzoxazepinone-Based WD Repeat Domain 5 Inhibitors.J Med Chem. 2023 Dec 28;66(24):16783-16806. doi: 10.1021/acs.jmedchem.3c01529. Epub 2023 Dec 12. J Med Chem. 2023. PMID: 38085679 Free PMC article.
-
WD repeat domain 5 (WDR5) inhibitors: a patent review (2016-present).Expert Opin Ther Pat. 2025 Jan;35(1):31-45. doi: 10.1080/13543776.2024.2441658. Epub 2024 Dec 22. Expert Opin Ther Pat. 2025. PMID: 39706200 Review.
-
The Development of Inhibitors Targeting the Mixed Lineage Leukemia 1 (MLL1)-WD Repeat Domain 5 Protein (WDR5) Protein- Protein Interaction.Curr Med Chem. 2020;27(33):5530-5542. doi: 10.2174/0929867326666190528080514. Curr Med Chem. 2020. PMID: 31132972 Review.
Cited by
-
Expanded profiling of WD repeat domain 5 inhibitors reveals actionable strategies for the treatment of hematologic malignancies.Proc Natl Acad Sci U S A. 2024 Aug 27;121(35):e2408889121. doi: 10.1073/pnas.2408889121. Epub 2024 Aug 21. Proc Natl Acad Sci U S A. 2024. PMID: 39167600 Free PMC article.
-
WDR5 Binding to Histone Serotonylation Is Driven by an Edge-Face Aromatic Interaction with Unexpected Electrostatic Effects.J Am Chem Soc. 2024 Oct 9;146(40):27451-27459. doi: 10.1021/jacs.4c07277. Epub 2024 Sep 25. J Am Chem Soc. 2024. PMID: 39321462
-
WDR5 represents a therapeutically exploitable target for cancer stem cells in glioblastoma.Genes Dev. 2023 Feb 1;37(3-4):86-102. doi: 10.1101/gad.349803.122. Epub 2023 Feb 2. Genes Dev. 2023. PMID: 36732025 Free PMC article.
-
Small molecule and peptide inhibitors of βTrCP and the βTrCP-NRF2 protein-protein interaction.Biochem Soc Trans. 2023 Jun 28;51(3):925-936. doi: 10.1042/BST20220352. Biochem Soc Trans. 2023. PMID: 37293994 Free PMC article. Review.
-
Structure-based discovery of potent WD repeat domain 5 inhibitors that demonstrate efficacy and safety in preclinical animal models.Proc Natl Acad Sci U S A. 2023 Jan 3;120(1):e2211297120. doi: 10.1073/pnas.2211297120. Epub 2022 Dec 27. Proc Natl Acad Sci U S A. 2023. PMID: 36574664 Free PMC article.
References
-
- Smith TF; Gaitatzes C; Saxena K; Neer EJ The WD Repeat: A Common Architecture for Diverse Functions. Trends Biochem. Sci. 1999, 24, 181–185. - PubMed
-
- Stirnimann CU; Petsalaki E; Russell RB; Müller, C. W. WD40 Proteins Propel Cellular Networks. Trends Biochem. Sci. 2010, 35, 565–574. - PubMed
-
- Couture JF; Collazo E; Trievel RC Molecular Recognition of Histone H3 by the WD40 Protein WDR5. Nat. Struct. Mol. Biol. 2006, 13, 698–703. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information
Molecular Biology Databases
Miscellaneous

