YqeH contributes to avian pathogenic Escherichia coli pathogenicity by regulating motility, biofilm formation, and virulence
- PMID: 35436977
- PMCID: PMC9014576
- DOI: 10.1186/s13567-022-01049-6
YqeH contributes to avian pathogenic Escherichia coli pathogenicity by regulating motility, biofilm formation, and virulence
Abstract
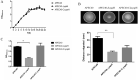
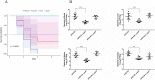
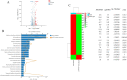
Avian pathogenic Escherichia coli (APEC) is a pathotype of extraintestinal pathogenic E. coli and one of the most serious infectious diseases of poultry. It not only causes great economic losses to the poultry industry, but also poses a serious threat to public health worldwide. Here, we examined the role of YqeH, a transcriptional regulator located at E. coli type III secretion system 2 (ETT2), in APEC pathogenesis. To investigate the effects of YqeH on APEC phenotype and virulence, we constructed a yqeH deletion mutant (APEC40-ΔyqeH) and a complemented strain (APEC40-CΔyqeH) of APEC40. Compared with the wild type (WT), the motility and biofilm formation of APEC40-ΔyqeH were significantly reduced. The yqeH mutant was highly attenuated in a chick infection model compared with WT, and showed severe defects in its adherence to and invasion of chicken embryo fibroblast DF-1 cells. However, the mechanisms underlying these phenomena were unclear. Therefore, we analyzed the transcriptional effects of the yqeH deletion to clarify the regulatory mechanisms of YqeH, and the role of YqeH in APEC virulence. The deletion of yqeH downregulated the transcript levels of several flagellum-, biofilm-, and virulence-related genes. Our results demonstrate that YqeH is involved in APEC pathogenesis, and the reduced virulence of APEC40-ΔyqeH may be related to its reduced motility and biofilm formation.
Keywords: Avian pathogenic Escherichia coli; biofilm formation; motility; pathogenicity; virulence; yqeH.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

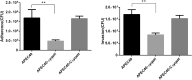
Similar articles
-
The Escherichia coli type III secretion system 2 Is involved in the biofilm formation and virulence of avian Pathogenic Escherichia coli.Comp Immunol Microbiol Infect Dis. 2021 Dec;79:101722. doi: 10.1016/j.cimid.2021.101722. Epub 2021 Nov 18. Comp Immunol Microbiol Infect Dis. 2021. PMID: 34823134
-
Flagellar rotor protein FliG is involved in the virulence of avian pathogenic Escherichia coli.Microb Pathog. 2021 Nov;160:105198. doi: 10.1016/j.micpath.2021.105198. Epub 2021 Sep 16. Microb Pathog. 2021. PMID: 34537273
-
LuxR family transcriptional repressor YjjQ modulates the biofilm formation and motility of avian pathogenic Escherichia coli.Res Vet Sci. 2022 Dec 20;152:10-19. doi: 10.1016/j.rvsc.2022.07.011. Epub 2022 Jul 22. Res Vet Sci. 2022. PMID: 35901637
-
Identification of novel biofilm genes in avian pathogenic Escherichia coli by Tn5 transposon mutant library.World J Microbiol Biotechnol. 2022 Jun 11;38(8):130. doi: 10.1007/s11274-022-03314-4. World J Microbiol Biotechnol. 2022. PMID: 35688968 Review.
-
[Avian pathogenic Escherichia coli (APEC)].Berl Munch Tierarztl Wochenschr. 2003 Sep-Oct;116(9-10):381-95. Berl Munch Tierarztl Wochenschr. 2003. PMID: 14526468 Review. German.
Cited by
-
FdeC expression regulates motility and adhesion of the avian pathogenic Escherichia coli strain IMT5155.Vet Res. 2024 May 31;55(1):70. doi: 10.1186/s13567-024-01327-5. Vet Res. 2024. PMID: 38822378 Free PMC article.
-
Characteristics, pathogenic mechanism, zoonotic potential, drug resistance, and prevention of avian pathogenic Escherichia coli (APEC).Front Microbiol. 2022 Dec 13;13:1049391. doi: 10.3389/fmicb.2022.1049391. eCollection 2022. Front Microbiol. 2022. PMID: 36583051 Free PMC article. Review.
-
Genetic distribution, characterization, and function of Escherichia coli type III secretion system 2 (ETT2).iScience. 2024 Apr 18;27(5):109763. doi: 10.1016/j.isci.2024.109763. eCollection 2024 May 17. iScience. 2024. PMID: 38706860 Free PMC article. Review.
-
EspE3 plays a role in the pathogenicity of avian pathogenic Escherichia coli.Vet Res. 2023 Aug 29;54(1):70. doi: 10.1186/s13567-023-01202-9. Vet Res. 2023. PMID: 37644523 Free PMC article.
References
-
- Apostolakos I, Laconi A, Mughini-Gras L, Yapicier Ö, Piccirillo A. Occurrence of colibacillosis in broilers and its relationship with avian pathogenic Escherichia coli (APEC) population structure and molecular characteristics. Front Vet Sci. 2021;8:737720. doi: 10.3389/fvets.2021.737720. - DOI - PMC - PubMed
-
- Paudel S, Fink D, Abdelhamid MK, Zöggeler A, Liebhart D, Hess M, Hess C. Aerosol is the optimal route of respiratory tract infection to induce pathological lesions of colibacillosis by a lux-tagged avian pathogenic Escherichia coli in chickens. Avian Pathol. 2021;50:417–426. doi: 10.1080/03079457.2021.1978392. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

