Breast tumor microenvironment structures are associated with genomic features and clinical outcome
- PMID: 35437329
- PMCID: PMC7612730
- DOI: 10.1038/s41588-022-01041-y
Breast tumor microenvironment structures are associated with genomic features and clinical outcome
Abstract
The functions of the tumor microenvironment (TME) are orchestrated by precise spatial organization of specialized cells, yet little is known about the multicellular structures that form within the TME. Here we systematically mapped TME structures in situ using imaging mass cytometry and multitiered spatial analysis of 693 breast tumors linked to genomic and clinical data. We identified ten recurrent TME structures that varied by vascular content, stromal quiescence versus activation, and leukocyte composition. These TME structures had distinct enrichment patterns among breast cancer subtypes, and some were associated with genomic profiles indicative of immune escape. Regulatory and dysfunctional T cells co-occurred in large 'suppressed expansion' structures. These structures were characterized by high cellular diversity, proliferating cells and enrichment for BRCA1 and CASP8 mutations and predicted poor outcome in estrogen-receptor-positive disease. The multicellular structures revealed here link conserved spatial organization to local TME function and could improve patient stratification.
© 2022. The Author(s).
Conflict of interest statement
C.C. is a member of AstraZeneca’s iMED External Science Panel and Illumina’s Scientific Advisory Board and a recipient of research grants (administered by the University of Cambridge) from Genentech, Roche, AstraZeneca and Servier. B.B. holds a patent relevant to this work entitled ‘A method for determining the likelihood of a patient being responsive to cancer immunotherapy’ (publication number WO2020207771A1). E.P. has received honoraria from Roche and Novartis for speaking at meetings and Inflection Point Biomedical Advisors for participating in an advisory panel. The other authors declare no competing interests.
Figures








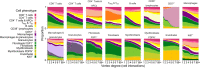


References
Publication types
MeSH terms
Grants and funding
- BRC-1215-20014/DH_/Department of Health/United Kingdom
- A7199/CRUK_/Cancer Research UK/United Kingdom
- 694620/ERC_/European Research Council/International
- A25815/CRUK_/Cancer Research UK/United Kingdom
- 25815/CRUK_/Cancer Research UK/United Kingdom
- MC_UU_00002/16/MRC_/Medical Research Council/United Kingdom
- A17197/CRUK_/Cancer Research UK/United Kingdom
- 16942/CRUK_/Cancer Research UK/United Kingdom
- 29567/CRUK_/Cancer Research UK/United Kingdom
- A29580/CRUK_/Cancer Research UK/United Kingdom
- NF-SI-0515-10090/DH_/Department of Health/United Kingdom
- A27657/CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

