This is a preprint.
Evolution of nasal and olfactory infection characteristics of SARS-CoV-2 variants
- PMID: 35441175
- PMCID: PMC9016639
- DOI: 10.1101/2022.04.12.487379
Evolution of nasal and olfactory infection characteristics of SARS-CoV-2 variants
Update in
-
Evolution of nasal and olfactory infection characteristics of SARS-CoV-2 variants.J Clin Invest. 2024 Mar 14;134(8):e174439. doi: 10.1172/JCI174439. J Clin Invest. 2024. PMID: 38483537 Free PMC article.
Abstract
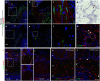
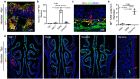
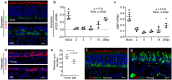
SARS-CoV-2 infection of the upper airway and the subsequent immune response are early, critical factors in COVID-19 pathogenesis. By studying infection of human biopsies in vitro and in a hamster model in vivo, we demonstrated a transition in tropism from olfactory to respiratory epithelium as the virus evolved. Analyzing each variants revealed that SARS-CoV-2 WA1 or Delta infects a proportion of olfactory neurons in addition to the primary target sustentacular cells. The Delta variant possesses broader cellular invasion capacity into the submucosa, while Omicron displays longer retention in the sinonasal epithelium. The olfactory neuronal infection by WA1 and the subsequent olfactory bulb transport via axon is more pronounced in younger hosts. In addition, the observed viral clearance delay and phagocytic dysfunction in aged olfactory mucosa is accompanied by a decline of phagocytosis related genes. Furthermore, robust basal stem cell activation contributes to neuroepithelial regeneration and restores ACE2 expression post-infection. Together, our study characterized the nasal tropism of SARS-CoV-2 strains, immune clearance, and regeneration post infection. The shifting characteristics of viral infection at the airway portal provides insight into the variability of COVID-19 clinical features and may suggest differing strategies for early local intervention.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures











References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
