Functional analysis of human olfactory receptors with a high basal activity using LNCaP cell line
- PMID: 35446888
- PMCID: PMC9022881
- DOI: 10.1371/journal.pone.0267356
Functional analysis of human olfactory receptors with a high basal activity using LNCaP cell line
Abstract
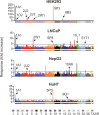
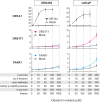
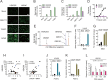
Humans use a family of more than 400 olfactory receptors (ORs) to detect odorants. However, deorphanization of ORs is a critical issue because the functional properties of more than 80% of ORs remain unknown, thus, hampering our understanding of the relationship between receptor function and perception. HEK293 cells are the most commonly used heterologous expression system to determine the function of a given OR; however, they cannot functionally express a majority of ORs probably due to a lack of factor(s) required in cells in which ORs function endogenously. Interestingly, ORs have been known to be expressed in a variety of cells outside the nose and play critical physiological roles. These findings prompted us to test the capacity of cells to functionally express a specific repertoire of ORs. In this study, we selected three cell lines that endogenously express functional ORs. We demonstrated that human prostate carcinoma (LNCaP) cell lines successfully identified novel ligands for ORs that were not recognized when expressed in HEK293 cells. Further experiments suggested that the LNCaP cell line was effective for functional expression of ORs, especially with a high basal activity, which impeded the sensitive detection of ligand-mediated activity of ORs. This report provides an efficient functional assay system for a specific repertoire of ORs that cannot be characterized in current cell systems.
Conflict of interest statement
All authors are employees of Kao Corporation. Kao Corporation has a patent (P6886893) related to an assay method for ORs expressed in LNCaP cell line. However, this does not alter the authors’ adherence to all PLOS ONE policies on data and material sharing. There are no products currently in development or marketed to declare.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

