Modular and mechanistic changes across stages of colorectal cancer
- PMID: 35448980
- PMCID: PMC9022252
- DOI: 10.1186/s12885-022-09479-3
Modular and mechanistic changes across stages of colorectal cancer
Abstract
Background: While mechanisms contributing to the progression and metastasis of colorectal cancer (CRC) are well studied, cancer stage-specific mechanisms have been less comprehensively explored. This is the focus of this manuscript.
Methods: Using previously published data for CRC (Gene Expression Omnibus ID GSE21510), we identified differentially expressed genes (DEGs) across four stages of the disease. We then generated unweighted and weighted correlation networks for each of the stages. Communities within these networks were detected using the Louvain algorithm and topologically and functionally compared across stages using the normalized mutual information (NMI) metric and pathway enrichment analysis, respectively. We also used Short Time-series Expression Miner (STEM) algorithm to detect potential biomarkers having a role in CRC.
Results: Sixteen Thousand Sixty Two DEGs were identified between various stages (p-value ≤ 0.05). Comparing communities of different stages revealed that neighboring stages were more similar to each other than non-neighboring stages, at both topological and functional levels. A functional analysis of 24 cancer-related pathways indicated that several signaling pathways were enriched across all stages. However, the stage-unique networks were distinctly enriched only for a subset of these 24 pathways (e.g., MAPK signaling pathway in stages I-III and Notch signaling pathway in stages III and IV). We identified potential biomarkers, including HOXB8 and WNT2 with increasing, and MTUS1 and SFRP2 with decreasing trends from stages I to IV. Extracting subnetworks of 10 cancer-relevant genes and their interacting first neighbors (162 genes in total) revealed that the connectivity patterns for these genes were different across stages. For example, BRAF and CDK4, members of the Ser/Thr kinase, up-regulated in cancer, displayed changing connectivity patterns from stages I to IV.
Conclusions: Here, we report molecular and modular networks for various stages of CRC, providing a pseudo-temporal view of the mechanistic changes associated with the disease. Our analysis highlighted similarities at both functional and topological levels, across stages. We further identified stage-specific mechanisms and biomarkers potentially contributing to the progression of CRC.
Keywords: Biomarkers; CRC stages; Colorectal cancer; Signaling pathways; Stage-specific networks; Stage-unique networks.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
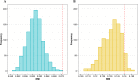
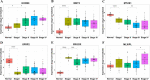
Figures




Similar articles
-
Identification of disrupted pathways in ulcerative colitis-related colorectal carcinoma by systematic tracking the dysregulated modules.J BUON. 2016 Mar-Apr;21(2):366-74. J BUON. 2016. PMID: 27273946
-
Identification of genes involved in the four stages of colorectal cancer: Gene expression profiling.Mol Cell Probes. 2018 Feb;37:39-47. doi: 10.1016/j.mcp.2017.11.004. Epub 2017 Nov 24. Mol Cell Probes. 2018. PMID: 29179987
-
Employing bioinformatics analysis to identify hub genes and microRNAs involved in colorectal cancer.Med Oncol. 2021 Aug 14;38(9):114. doi: 10.1007/s12032-021-01543-5. Med Oncol. 2021. PMID: 34390411
-
Identification of potential therapeutic targets associated with diagnosis and prognosis of colorectal cancer patients based on integrated bioinformatics analysis.Comput Biol Med. 2022 Jul;146:105688. doi: 10.1016/j.compbiomed.2022.105688. Epub 2022 May 31. Comput Biol Med. 2022. PMID: 35680454
-
Identification of MFI2-AS1, a Novel Pivotal lncRNA for Prognosis of Stage III/IV Colorectal Cancer.Dig Dis Sci. 2020 Dec;65(12):3538-3550. doi: 10.1007/s10620-020-06064-1. Epub 2020 Jan 20. Dig Dis Sci. 2020. PMID: 31960204 Review.
Cited by
-
FUT2 promotes the tumorigenicity and metastasis of colorectal cancer cells via the Wnt/β‑catenin pathway.Int J Oncol. 2023 Mar;62(3):35. doi: 10.3892/ijo.2023.5483. Epub 2023 Feb 3. Int J Oncol. 2023. PMID: 36734282 Free PMC article.
-
COADREADx: A comprehensive algorithmic dissection of colorectal cancer unravels salient biomarkers and actionable insights into its discrete progression.PeerJ. 2024 Oct 28;12:e18347. doi: 10.7717/peerj.18347. eCollection 2024. PeerJ. 2024. PMID: 39484215 Free PMC article.
-
Colorectal cancer with low SLC35A3 is associated with immune infiltrates and poor prognosis.Sci Rep. 2024 Jan 3;14(1):329. doi: 10.1038/s41598-023-51028-w. Sci Rep. 2024. PMID: 38172565 Free PMC article.
-
Development and validation of survival prediction tools in early and late onset colorectal cancer patients.Sci Rep. 2025 Apr 15;15(1):12864. doi: 10.1038/s41598-025-95385-0. Sci Rep. 2025. PMID: 40229310 Free PMC article.
-
Polyphenol Intake in Elderly Patients: A Novel Approach to Counteract Colorectal Cancer Risk?Int J Mol Sci. 2025 Mar 11;26(6):2497. doi: 10.3390/ijms26062497. Int J Mol Sci. 2025. PMID: 40141143 Free PMC article. Review.
References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J Clinicians. 2018;68(6):394–424. - PubMed
-
- Pawa N, Arulampalam T, Norton JD. Screening for colorectal cancer: established and emerging modalities. Nat Rev Gastroenterol Hepatol. 2011;8(12):711–722. - PubMed
MeSH terms
Substances
Grants and funding
- U01 DK097430/DK/NIDDK NIH HHS/United States
- R01 HL106579/HL/NHLBI NIH HHS/United States
- U01 DK097430, U01 CA200147, U01 CA198941, U19 AI090023, R01 HL106579, R01HL108735, R01 HD084633, R01 DK109365, R01LM012595, OTA OD030544/NH/NIH HHS/United States
- R01 LM012595/LM/NLM NIH HHS/United States
- OT2 OD030544/OD/NIH HHS/United States
- R01 DK109365/DK/NIDDK NIH HHS/United States
- R01 HD084633/HD/NICHD NIH HHS/United States
- U01 CA200147/CA/NCI NIH HHS/United States
- R01 HL085159/HL/NHLBI NIH HHS/United States
- U01 CA198941/CA/NCI NIH HHS/United States
- U19 AI090023/AI/NIAID NIH HHS/United States
- R01 HL108735/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials

