R-loops at microRNA encoding loci promote co-transcriptional processing of pri-miRNAs in plants
- PMID: 35449404
- PMCID: PMC9023350
- DOI: 10.1038/s41477-022-01125-x
R-loops at microRNA encoding loci promote co-transcriptional processing of pri-miRNAs in plants
Abstract
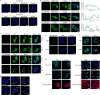
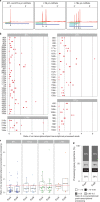
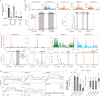
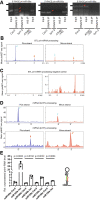
In most organisms, the maturation of nascent RNAs is coupled to transcription. Unlike in animals, the RNA polymerase II (RNAPII) transcribes microRNA genes (MIRNAs) as long and structurally variable pri-miRNAs in plants. Current evidence suggests that the miRNA biogenesis complex assembly initiates early during the transcription of pri-miRNAs in plants. However, it is unknown whether miRNA processing occurs co-transcriptionally. Here, we used native elongating transcript sequencing data and imaging techniques to demonstrate that plant miRNA biogenesis occurs coupled to transcription. We found that the entire biogenesis occurs co-transcriptionally for pri-miRNAs processed from the loop of the hairpin but requires a second nucleoplasmic step for those processed from the base. Furthermore, we found that co- and post-transcriptional miRNA processing mechanisms co-exist for most miRNAs in a dynamic balance. Notably, we discovered that R-loops, formed near the transcription start site region of MIRNAs, promote co-transcriptional pri-miRNA processing. Furthermore, our results suggest the neofunctionalization of co-transcriptionally processed miRNAs, boosting countless regulatory scenarios.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






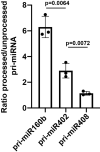

Comment in
-
The R-loop influences miRNA birth place.Nat Plants. 2022 Apr;8(4):320-321. doi: 10.1038/s41477-022-01124-y. Nat Plants. 2022. PMID: 35449403 No abstract available.
Similar articles
-
Primary microRNA processing is functionally coupled to RNAP II transcription in vitro.Sci Rep. 2015 Jul 7;5:11992. doi: 10.1038/srep11992. Sci Rep. 2015. PMID: 26149087 Free PMC article.
-
Identification of Pri-miRNA Stem-Loop Interacting Proteins in Plants Using a Modified Version of the Csy4 CRISPR Endonuclease.Int J Mol Sci. 2022 Aug 11;23(16):8961. doi: 10.3390/ijms23168961. Int J Mol Sci. 2022. PMID: 36012225 Free PMC article.
-
Microprocessor Recruitment to Elongating RNA Polymerase II Is Required for Differential Expression of MicroRNAs.Cell Rep. 2017 Sep 26;20(13):3123-3134. doi: 10.1016/j.celrep.2017.09.010. Cell Rep. 2017. PMID: 28954229 Free PMC article.
-
Recent advances in the regulation of plant miRNA biogenesis.RNA Biol. 2021 Dec;18(12):2087-2096. doi: 10.1080/15476286.2021.1899491. Epub 2021 Mar 17. RNA Biol. 2021. PMID: 33666136 Free PMC article. Review.
-
New insights into pri-miRNA processing and accumulation in plants.Wiley Interdiscip Rev RNA. 2015 Sep-Oct;6(5):533-45. doi: 10.1002/wrna.1292. Epub 2015 Jun 29. Wiley Interdiscip Rev RNA. 2015. PMID: 26119101 Review.
Cited by
-
Exploring the landscape of miRNA production and the structural rules that shape it.Nat Plants. 2024 Jul;10(7):1060-1061. doi: 10.1038/s41477-024-01728-6. Nat Plants. 2024. PMID: 38956418 No abstract available.
-
Cotranscriptional RNA processing and modification in plants.Plant Cell. 2023 May 29;35(6):1654-1670. doi: 10.1093/plcell/koac309. Plant Cell. 2023. PMID: 36259932 Free PMC article. Review.
-
The expanding role of RNA modifications in plant RNA polymerase II transcripts: highlights and perspectives.J Exp Bot. 2023 Aug 3;74(14):3975-3986. doi: 10.1093/jxb/erad136. J Exp Bot. 2023. PMID: 37076273 Free PMC article. Review.
-
Intrinsically disordered proteins SAID1/2 condensate on SERRATE for dual inhibition of miRNA biogenesis in Arabidopsis.Proc Natl Acad Sci U S A. 2023 Apr 4;120(14):e2216006120. doi: 10.1073/pnas.2216006120. Epub 2023 Mar 27. Proc Natl Acad Sci U S A. 2023. PMID: 36972460 Free PMC article.
-
A plant bunyaviral protein disrupts SERRATE phase separation to modulate microRNA biogenesis during viral pathogenesis.Nat Commun. 2025 Jul 4;16(1):6197. doi: 10.1038/s41467-025-61528-0. Nat Commun. 2025. PMID: 40615422 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

