Histone Deacetylase Inhibitors Impair Glioblastoma Cell Motility and Proliferation
- PMID: 35454804
- PMCID: PMC9027190
- DOI: 10.3390/cancers14081897
Histone Deacetylase Inhibitors Impair Glioblastoma Cell Motility and Proliferation
Abstract
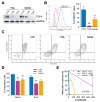
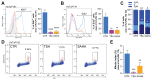
Despite being subjected to high-dose chemo and radiotherapy, glioblastoma (GBM) patients still encounter almost inevitable relapse, due to the capability of tumor cells to disseminate and invade normal brain tissues. Moreover, the presence of a cancer stem cell (CSC) subpopulation, already demonstrated to better resist and evade treatments, further frustrates potential therapeutic approaches. In this context, we previously demonstrated that GBM is characterized by a tightly-regulated balance between the β-catenin cofactors TCF1 and TCF4, with high levels of TCF4 responsible for sustaining CSC in these tumors; thus, supporting their aggressive features. Since histone deacetylase inhibitors (HDI) have been reported to strongly reduce TCF4 levels in colon cancer cells, we hypothesized that they could also exert a similar therapeutic action in GBM. Here, we treated primary GBM cultures with Trichostatin-A and Vorinostat, demonstrating their ability to strongly suppress the Wnt-dependent pathways; thus, promoting CSC differentiation and concomitantly impairing GBM cell viability and proliferation. More interestingly, analysis of their molecular effects suggested a prominent HDI action against GBM cell motility/migration, which we demonstrated to rely on the inhibition of the RhoA-GTPase and interferon intracellular cascades. Our results suggest HDI as potential therapeutic agents in GBM, through their action on multiple cancer hallmarks.
Keywords: Wnt signaling; cell migration; glioblastoma; histone deacetylase inhibitors; interferon pathway.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Verhaak R.G., Hoadley K.A., Purdom E., Wang V., Qi Y., Wilkerson M.D., Miller C.R., Ding L., Golub T., Mesirov J.P., et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17:98–110. doi: 10.1016/j.ccr.2009.12.020. - DOI - PMC - PubMed
-
- Marenco-Hillembrand L., Wijesekera O., Suarez-Meade P., Mampre D., Jackson C., Peterson J., Trifiletti D., Hammack J., Ortiz K., Lesser E., et al. Trends in glioblastoma: Outcomes over time and type of intervention: A systematic evidence based analysis. J. Neurooncol. 2020;147:297–307. doi: 10.1007/s11060-020-03451-6. - DOI - PubMed
-
- Stupp R., Hegi M.E., Mason W.P., van den Bent M.J., Taphoorn M.J., Janzer R.C., Ludwin S.K., Allgeier A., Fisher B., Belanger K., et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 2009;10:459–466. doi: 10.1016/S1470-2045(09)70025-7. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

