Population Genetic Structure and Hybridization of Schistosoma haematobium in Nigeria
- PMID: 35456103
- PMCID: PMC9026724
- DOI: 10.3390/pathogens11040425
Population Genetic Structure and Hybridization of Schistosoma haematobium in Nigeria
Abstract
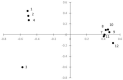
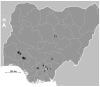
Background: Schistosomiasis is a major poverty-related disease caused by dioecious parasitic flatworms of the genus Schistosoma with a health impact on both humans and animals. Hybrids of human urogenital schistosome and bovine intestinal schistosome have been reported in humans in several of Nigeria’s neighboring West African countries. No empirical studies have been carried out on the genomic diversity of Schistosoma haematobium in Nigeria. Here, we present novel data on the presence and prevalence of hybrids and the population genetic structure of S. haematobium. Methods: 165 Schistosoma-positive urine samples were obtained from 12 sampling sites in Nigeria. Schistosoma haematobium eggs from each sample were hatched and each individual miracidium was picked and preserved in Whatman® FTA cards for genomic analysis. Approximately 1364 parasites were molecularly characterized by rapid diagnostic multiplex polymerase chain reaction (RD-PCR) for mitochondrial DNA gene (Cox1 mtDNA) and a subset of 1136 miracidia were genotyped using a panel of 18 microsatellite markers. Results: No significant difference was observed in the population genetic diversity (p > 0.05), though a significant difference was observed in the allelic richness of the sites except sites 7, 8, and 9 (p < 0.05). Moreover, we observed two clusters of populations: west (populations 1−4) and east (populations 7−12). Of the 1364 miracidia genotyped, 1212 (89%) showed an S. bovis Cox1 profile and 152 (11%) showed an S. haematobium cox1 profile. All parasites showed an S. bovis Cox1 profile except for some at sites 3 and 4. Schistosoma miracidia full genotyping showed 59.3% of the S. bovis ITS2 allele. Conclusions: This study provides novel insight into hybridization and population genetic structure of S. haematobium in Nigeria. Our findings suggest that S. haematobium x S. bovis hybrids are common in Nigeria. More genomic studies on both human- and animal-infecting parasites are needed to ascertain the role of animals in schistosome transmission.
Keywords: Nigeria; Schistosoma bovis; Schistosoma haematobium; hybrids; population genetic analysis.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of manuscript, or in the decision to publish the results.
Figures

Similar articles
-
Population genetic structure of Schistosoma haematobium and Schistosoma haematobium × Schistosoma bovis hybrids among school-aged children in Côte d'Ivoire.Parasite. 2022;29:23. doi: 10.1051/parasite/2022023. Epub 2022 May 3. Parasite. 2022. PMID: 35522066 Free PMC article.
-
Hybridization increases genetic diversity in Schistosoma haematobium populations infecting humans in Cameroon.Infect Dis Poverty. 2022 Mar 26;11(1):37. doi: 10.1186/s40249-022-00958-0. Infect Dis Poverty. 2022. PMID: 35346375 Free PMC article.
-
High prevalence of Schistosoma haematobium × Schistosoma bovis hybrids in schoolchildren in Côte d'Ivoire.Parasitology. 2020 Mar;147(3):287-294. doi: 10.1017/S0031182019001549. Epub 2019 Nov 21. Parasitology. 2020. PMID: 31727202 Free PMC article.
-
Mapping of schistosome hybrids of the haematobium group in West and Central Africa.J Helminthol. 2024 Sep 18;98:e53. doi: 10.1017/S0022149X24000257. J Helminthol. 2024. PMID: 39291545 Review.
-
Natural Intra- and Interclade Human Hybrid Schistosomes in Africa with Considerations on Prevention through Vaccination.Microorganisms. 2021 Jul 8;9(7):1465. doi: 10.3390/microorganisms9071465. Microorganisms. 2021. PMID: 34361901 Free PMC article. Review.
Cited by
-
Human-water interactions associated to cercarial emergence pattern and their influences on urinary schistosomiasis transmission in two endemic areas in Mali.Infect Dis Poverty. 2024 Aug 29;13(1):62. doi: 10.1186/s40249-024-01229-w. Infect Dis Poverty. 2024. PMID: 39198901 Free PMC article.
-
The impact of Schistosoma haematobium hybridization on molecular diagnosis of schistosomiasis: A review with emphasis on female genital schistosomiasis.PLoS Negl Trop Dis. 2025 Aug 7;19(8):e0013364. doi: 10.1371/journal.pntd.0013364. eCollection 2025 Aug. PLoS Negl Trop Dis. 2025. PMID: 40773459 Free PMC article. Review.
-
Imported Schistosomiasis in Southwestern Europe: Wide Variation of Pure and Hybrid Genotypes Infecting Sub-Saharan Migrants.Transbound Emerg Dis. 2025 Apr 18;2025:6614509. doi: 10.1155/tbed/6614509. eCollection 2025. Transbound Emerg Dis. 2025. PMID: 40718473 Free PMC article.
-
Reaching the World Health Organization elimination targets for schistosomiasis: the importance of a One Health perspective.Philos Trans R Soc Lond B Biol Sci. 2023 Oct 9;378(1887):20220274. doi: 10.1098/rstb.2022.0274. Epub 2023 Aug 21. Philos Trans R Soc Lond B Biol Sci. 2023. PMID: 37598697 Free PMC article. Review.
-
Genetic characterization of schistosome species from cattle in Côte d'Ivoire.Parasit Vectors. 2024 Mar 12;17(1):122. doi: 10.1186/s13071-024-06221-9. Parasit Vectors. 2024. PMID: 38475876 Free PMC article.
References
-
- Hotez P.J., Alvarado M., Basanez M.G., Bolliger I., Bourne R., Boussinesq M., Brooker S.J., Brown A.S., Buckle G., Budke C.M., et al. The global burden of disease study 2010: Interpretation and implications for the neglected tropical diseases. PLoS Negl. Trop Dis. 2014;8:e2865. doi: 10.1371/journal.pntd.0002865. - DOI - PMC - PubMed
-
- Boissier J., Mouahid G., Moné H. In: Schistosoma spp. (Robertson, L (eds) Part 4 Helminths) Rose J.B., Jiménez-Cisneros B., editors. Michigan State University; E. Lansing, MI, USA: 2019.
-
- Mone H., Mouahid G., Morand S. The distribution of Schistosoma bovis Sonsino, 1876 in relation to intermediate host mollusc-parasite relationships. Adv. Parasitol. 1999;44:99–138. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

