Transcriptomic Analysis of Canine Osteosarcoma from a Precision Medicine Perspective Reveals Limitations of Differential Gene Expression Studies
- PMID: 35456486
- PMCID: PMC9031617
- DOI: 10.3390/genes13040680
Transcriptomic Analysis of Canine Osteosarcoma from a Precision Medicine Perspective Reveals Limitations of Differential Gene Expression Studies
Abstract
Despite significant advances in cancer diagnosis and treatment, osteosarcoma (OSA), an aggressive primary bone tumor, has eluded attempts at improving patient survival for many decades. The difficulty in managing OSA lies in its extreme genetic complexity, drug resistance, and heterogeneity, making it improbable that a single-target treatment would be beneficial for the majority of affected individuals. Precision medicine seeks to fill this gap by addressing the intra- and inter-tumoral heterogeneity to improve patient outcome and survival. The characterization of differentially expressed genes (DEGs) unique to the tumor provides insight into the phenotype and can be useful for informing appropriate therapies as well as the development of novel treatments. Traditional DEG analysis combines patient data to derive statistically inferred genes that are dysregulated in the group; however, the results from this approach are not necessarily consistent across individual patients, thus contradicting the basis of precision medicine. Spontaneously occurring OSA in the dog shares remarkably similar clinical, histological, and molecular characteristics to the human disease and therefore serves as an excellent model. In this study, we use transcriptomic sequencing of RNA isolated from primary OSA tumor and patient-matched normal bone from seven dogs prior to chemotherapy to identify DEGs in the group. We then evaluate the universality of these changes in transcript levels across patients to identify DEGs at the individual level. These results can be useful for reframing our perspective of transcriptomic analysis from a precision medicine perspective by identifying variations in DEGs among individuals.
Keywords: cancer; canine; osteosarcoma; sequencing; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

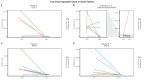
References
-
- Geller D.S., Gorlick R. Osteosarcoma: A Review of Diagnosis, Management, and Treatment Strategies. Clin. Adv. Hematol. Oncol. 2010;8:705–718. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

