Molecular Dynamics and Evolution of Centromeres in the Genus Equus
- PMID: 35457002
- PMCID: PMC9024551
- DOI: 10.3390/ijms23084183
Molecular Dynamics and Evolution of Centromeres in the Genus Equus
Abstract
The centromere is the chromosomal locus essential for proper chromosome segregation. While the centromeric function is well conserved and epigenetically specified, centromeric DNA sequences are typically composed of satellite DNA and represent the most rapidly evolving sequences in eukaryotic genomes. The presence of satellite sequences at centromeres hampered the comprehensive molecular analysis of these enigmatic loci. The discovery of functional centromeres completely devoid of satellite repetitions and fixed in some animal and plant species represented a turning point in centromere biology, definitively proving the epigenetic nature of the centromere. The first satellite-free centromere, fixed in a vertebrate species, was discovered in the horse. Later, an extraordinary number of satellite-free neocentromeres had been discovered in other species of the genus Equus, which remains the only mammalian genus with numerous satellite-free centromeres described thus far. These neocentromeres arose recently during evolution and are caught in a stage of incomplete maturation. Their presence made the equids a unique model for investigating, at molecular level, the minimal requirements for centromere seeding and evolution. This model system provided new insights on how centromeres are established and transmitted to the progeny and on the role of satellite DNA in different aspects of centromere biology.
Keywords: CENP-A; centromere repositioning; centromere sliding; genus Equus; karyotype evolution; neocentromeres; satellite DNA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
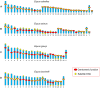
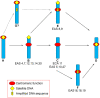
References
Publication types
MeSH terms
Substances
Grants and funding
- Dipartimenti di Eccellenza Program (2018-2022)-Dept. of Biology and Biotechnology "L. Spallanzani", University of Pavia)/Ministry of Education, Universities and Research
- PRIN Grant No. 2015RA7XZS_002/Ministry of Education, Universities and Research
- Progetto Bandiera Epigenomica/National Research Council
LinkOut - more resources
Full Text Sources

