Validation of artificial intelligence prediction models for skin cancer diagnosis using dermoscopy images: the 2019 International Skin Imaging Collaboration Grand Challenge
- PMID: 35461690
- PMCID: PMC9295694
- DOI: 10.1016/S2589-7500(22)00021-8
Validation of artificial intelligence prediction models for skin cancer diagnosis using dermoscopy images: the 2019 International Skin Imaging Collaboration Grand Challenge
Abstract
Background: Previous studies of artificial intelligence (AI) applied to dermatology have shown AI to have higher diagnostic classification accuracy than expert dermatologists; however, these studies did not adequately assess clinically realistic scenarios, such as how AI systems behave when presented with images of disease categories that are not included in the training dataset or images drawn from statistical distributions with significant shifts from training distributions. We aimed to simulate these real-world scenarios and evaluate the effects of image source institution, diagnoses outside of the training set, and other image artifacts on classification accuracy, with the goal of informing clinicians and regulatory agencies about safety and real-world accuracy.
Methods: We designed a large dermoscopic image classification challenge to quantify the performance of machine learning algorithms for the task of skin cancer classification from dermoscopic images, and how this performance is affected by shifts in statistical distributions of data, disease categories not represented in training datasets, and imaging or lesion artifacts. Factors that might be beneficial to performance, such as clinical metadata and external training data collected by challenge participants, were also evaluated. 25 331 training images collected from two datasets (in Vienna [HAM10000] and Barcelona [BCN20000]) between Jan 1, 2000, and Dec 31, 2018, across eight skin diseases, were provided to challenge participants to design appropriate algorithms. The trained algorithms were then tested for balanced accuracy against the HAM10000 and BCN20000 test datasets and data from countries not included in the training dataset (Turkey, New Zealand, Sweden, and Argentina). Test datasets contained images of all diagnostic categories available in training plus other diagnoses not included in training data (not trained category). We compared the performance of the algorithms against that of 18 dermatologists in a simulated setting that reflected intended clinical use.
Findings: 64 teams submitted 129 state-of-the-art algorithm predictions on a test set of 8238 images. The best performing algorithm achieved 58·8% balanced accuracy on the BCN20000 data, which was designed to better reflect realistic clinical scenarios, compared with 82·0% balanced accuracy on HAM10000, which was used in a previously published benchmark. Shifted statistical distributions and disease categories not included in training data contributed to decreases in accuracy. Image artifacts, including hair, pen markings, ulceration, and imaging source institution, decreased accuracy in a complex manner that varied based on the underlying diagnosis. When comparing algorithms to expert dermatologists (2460 ratings on 1269 images), algorithms performed better than experts in most categories, except for actinic keratoses (similar accuracy on average) and images from categories not included in training data (26% correct for experts vs 6% correct for algorithms, p<0·0001). For the top 25 submitted algorithms, 47·1% of the images from categories not included in training data were misclassified as malignant diagnoses, which would lead to a substantial number of unnecessary biopsies if current state-of-the-art AI technologies were clinically deployed.
Interpretation: We have identified specific deficiencies and safety issues in AI diagnostic systems for skin cancer that should be addressed in future diagnostic evaluation protocols to improve safety and reliability in clinical practice.
Funding: Melanoma Research Alliance and La Marató de TV3.
Copyright © 2022 The Author(s). Published by Elsevier Ltd. This is an Open Access article under the CC BY-NC-ND 4.0 license. Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of interests NC was an employee of IBM during a portion of this work. IBM's only involvement was through the work of NC. NC is currently employed at Microsoft, but Microsoft's only involvement is through the work of NC. NC holds two dermatology patents (US patent 10568695 B2 for surgical skin lesion removal and US patent 10255674 B2 for surface reflectance reduction in images using non-specular portion replacement) that are not relevant to this work. NC reports holding stock in IBM and Microsoft; payment or honoraria from Memorial Sloan-Kettering; and support for attending meetings or travel from Memorial Sloan-Kettering and IBM. SPu consults or receives honoraria from Almirall, ISDIN, Bristol Myers Squibb, La Roche-Posay, Pfizer, Regeneron, Sanofi, and SunPharma. VR is an expert advisor for Inhabit Brands. PT receives honoraria from Silverchair and Lilly. BH is employed by Kitware. AH consults for Canfield Scientific. All other authors declare no competing interests.
Figures
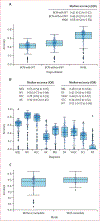




References
-
- Cancer Today. Population fact sheets. https://gco.iarc.fr/today/fact-sheets-populations (accessed July 7, 2020).
-
- Vestergaard ME, Macaskill P, Holt PE, Menzies SW. Dermoscopy compared with naked eye examination for the diagnosis of primary melanoma: a meta-analysis of studies performed in a clinical setting. Br J Dermatol 2008; 159: 669–76. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

