A Multi-Level Iterative Bi-Clustering Method for Discovering miRNA Co-regulation Network of Abiotic Stress Tolerance in Soybeans
- PMID: 35463453
- PMCID: PMC9021755
- DOI: 10.3389/fpls.2022.860791
A Multi-Level Iterative Bi-Clustering Method for Discovering miRNA Co-regulation Network of Abiotic Stress Tolerance in Soybeans
Abstract
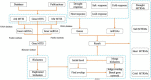
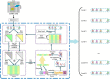
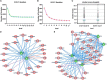
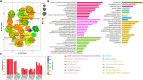
Although growing evidence shows that microRNA (miRNA) regulates plant growth and development, miRNA regulatory networks in plants are not well understood. Current experimental studies cannot characterize miRNA regulatory networks on a large scale. This information gap provides an excellent opportunity to employ computational methods for global analysis and generate valuable models and hypotheses. To address this opportunity, we collected miRNA-target interactions (MTIs) and used MTIs from Arabidopsis thaliana and Medicago truncatula to predict homologous MTIs in soybeans, resulting in 80,235 soybean MTIs in total. A multi-level iterative bi-clustering method was developed to identify 483 soybean miRNA-target regulatory modules (MTRMs). Furthermore, we collected soybean miRNA expression data and corresponding gene expression data in response to abiotic stresses. By clustering these data, 37 MTRMs related to abiotic stresses were identified, including stress-specific MTRMs and shared MTRMs. These MTRMs have gene ontology (GO) enrichment in resistance response, iron transport, positive growth regulation, etc. Our study predicts soybean MTRMs and miRNA-GO networks under different stresses, and provides miRNA targeting hypotheses for experimental analyses. The method can be applied to other biological processes and other plants to elucidate miRNA co-regulation mechanisms.
Keywords: abiotic stress tolerance in soybeans; bi-clustering; homology expansion; miRNA co-regulation; miRNA–target.
Copyright © 2022 Chang, Zhang, Zhang, Su, Qin, Li, Li, Wang, Zhao, Zhao, Zhao, Liu, Stacey and Xu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

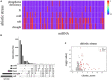

References
-
- Balyan S. C., Mutum R. D., Kansal S., Kumar S., Mathur S., Raghuvanshi S. (2015). “Insights into the small RNA-mediated networks in response to abiotic stress in plants,” in Elucidation of Abiotic Stress Signaling in Plants: Functional Genomics Gerspectives, Vol. 2 ed. Pandey G. K. (New York, NY: Springer; ), 45–92. 10.1007/978-1-4939-2540-7_3 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

