Comparison of miRNA and mRNA Expression in Sika Deer Testes With Age
- PMID: 35464385
- PMCID: PMC9019638
- DOI: 10.3389/fvets.2022.854503
Comparison of miRNA and mRNA Expression in Sika Deer Testes With Age
Abstract
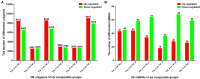
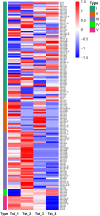
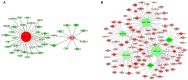
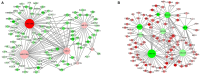
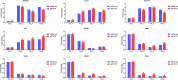
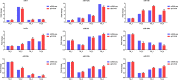
To elucidate the complex physiological process of testis development and spermatogenesis in Sika deer, this study evaluated the changes of miRNA and mRNA profiles in the four developmental stages of testis in the juvenile (1-year-old), adolescence (3-year-old), adult (5-year-old), and aged (10-year-old) stages. The results showed that a total of 198 mature, 66 novel miRNAs, and 23,558 differentially expressed (DE) unigenes were obtained; 14,918 (8,413 up and 6,505 down), 4,988 (2,453 up and 2,535 down), and 5,681 (2,929 up and 2,752 down) DE unigenes, as well as 88 (43 up and 45 down), 102 (44 up and 58 down), and 54 (18 up and 36 down) DE miRNAs were identified in 3- vs. 1-, 5- vs. 3-, and 10- vs. 5-year-old testes, respectively. By integrating miRNA and mRNA expression profiles, we predicted 10,790 mRNA-mRNA and 69,883 miRNA-mRNA interaction sites. The target genes were enriched by GO and KEGG pathways to obtain DE mRNA (IGF1R, ALKBH5, Piwil, HIF1A, BRDT, etc.) and DE miRNA (miR-140, miR-145, miR-7, miR-26a, etc.), which play an important role in testis development and spermatogenesis. The data show that DE miRNAs could regulate testis developmental and spermatogenesis through signaling pathways, including the MAPK signaling pathway, p53 signaling pathway, PI3K-Akt signaling pathway, Hippo signaling pathway, etc. miR-140 was confirmed to directly target mutant IGF1R-3'UTR by the Luciferase reporter assays. This study provides a useful resource for future studies on the role of miRNA regulation in testis development and spermatogenesis.
Keywords: mRNA; miRNA; sika deer; spermatogenesis; testis development.
Copyright © 2022 Jia, Zhang, Ma, Wang, Li, Diao, Leng, Shi, Zeng, Zong, Liu, Gong, Cai, Yang, Du and Chang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Altered miRNA and mRNA Expression in Sika Deer Skeletal Muscle with Age.Genes (Basel). 2020 Feb 6;11(2):172. doi: 10.3390/genes11020172. Genes (Basel). 2020. PMID: 32041309 Free PMC article.
-
Integrated Analysis of miRNA and mRNA Expression Profiles Reveals Functional miRNA-Targets in Development Testes of Small Tail Han Sheep.G3 (Bethesda). 2019 Feb 7;9(2):523-533. doi: 10.1534/g3.118.200947. G3 (Bethesda). 2019. PMID: 30559255 Free PMC article.
-
Integrated analysis of miRNA and mRNA expression profiles in testes of Duroc and Meishan boars.BMC Genomics. 2020 Oct 2;21(1):686. doi: 10.1186/s12864-020-07096-7. BMC Genomics. 2020. PMID: 33008286 Free PMC article.
-
Integrated analysis of miRNA and mRNA transcriptomic reveals antler growth regulatory network.Mol Genet Genomics. 2021 May;296(3):689-703. doi: 10.1007/s00438-021-01776-z. Epub 2021 Mar 26. Mol Genet Genomics. 2021. PMID: 33770271
-
Integrated analysis of miRNA and mRNA expression profiles in testes of Landrace and Hezuo boars.Front Vet Sci. 2022 Oct 18;9:942669. doi: 10.3389/fvets.2022.942669. eCollection 2022. Front Vet Sci. 2022. PMID: 36330159 Free PMC article.
Cited by
-
Hypoxia-induced m6A demethylase ALKBH5 promotes ovarian cancer tumorigenicity by decreasing methylation of the lncRNA RMRP.Am J Cancer Res. 2023 Sep 15;13(9):4179-4191. eCollection 2023. Am J Cancer Res. 2023. PMID: 37818080 Free PMC article.
-
The combination of SMRT sequencing and Illumina sequencing highlights organ-specific and age-specific expression patterns of miRNAs in Sika Deer.Front Vet Sci. 2022 Nov 14;9:1042445. doi: 10.3389/fvets.2022.1042445. eCollection 2022. Front Vet Sci. 2022. PMID: 36452144 Free PMC article.
-
Association between Sperm Morphology and Altered Sperm microRNA Expression.Biology (Basel). 2022 Nov 17;11(11):1671. doi: 10.3390/biology11111671. Biology (Basel). 2022. PMID: 36421385 Free PMC article.
-
Small RNA perspective of physical exercise-related improvement of male reproductive dysfunction due to obesity.Front Endocrinol (Lausanne). 2022 Dec 2;13:1038449. doi: 10.3389/fendo.2022.1038449. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 36531465 Free PMC article.
-
Association between Yili goose sperm motility and expression profiles of mRNA and miRNA in testis.BMC Genomics. 2023 Oct 24;24(1):640. doi: 10.1186/s12864-023-09727-1. BMC Genomics. 2023. PMID: 37875805 Free PMC article.
References
-
- Koziol K, Broda D, Romerowicz-Misielak M, Nowak S, Koziorowski M. Melatonin concentration in peripheral blood and melatonin receptors (MT1 and MT2) in the testis and epididymis of male roe deer during active spermatogenesis. Theriogenology. (2020) 149:25–37. 10.1016/j.theriogenology.2020.03.025 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

