Structure of a Vaccine-Induced, Germline-Encoded Human Antibody Defines a Neutralizing Epitope on the SARS-CoV-2 Spike N-Terminal Domain
- PMID: 35467422
- PMCID: PMC9239078
- DOI: 10.1128/mbio.03580-21
Structure of a Vaccine-Induced, Germline-Encoded Human Antibody Defines a Neutralizing Epitope on the SARS-CoV-2 Spike N-Terminal Domain
Abstract
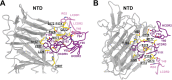
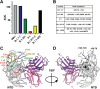
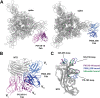
Structural characterization of infection- and vaccination-elicited antibodies in complex with antigen provides insight into the evolutionary arms race between the host and the pathogen and informs rational vaccine immunogen design. We isolated a germ line-encoded monoclonal antibody (mAb) from plasmablasts activated upon mRNA vaccination against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and determined its structure in complex with the spike glycoprotein by electron cryomicroscopy (cryo-EM). We show that the mAb engages a previously uncharacterized neutralizing epitope on the spike N-terminal domain (NTD). The high-resolution structure reveals details of the intermolecular interactions and shows that the mAb inserts its heavy complementarity-determining region 3 (HCDR3) loop into a hydrophobic NTD cavity previously shown to bind a heme metabolite, biliverdin. We demonstrate direct competition with biliverdin and that, because of the conserved nature of the epitope, the mAb maintains binding to viral variants B.1.1.7 (alpha), B.1.351 (beta), B.1.617.2 (delta), and B.1.1.529 (omicron). Our study describes a novel conserved epitope on the NTD that is readily targeted by vaccine-induced antibody responses. IMPORTANCE We report the first structure of a vaccine-induced antibody to SARS-CoV-2 spike isolated from plasmablasts 7 days after vaccination. The genetic sequence of the antibody PVI.V6-14 suggests that it is completely unmutated, meaning that this type of B cell did not undergo somatic hypermutation or affinity maturation; this cell was likely already present in the donor and was activated by the vaccine. This is, to our knowledge, also the first structure of an unmutated antibody in complex with its cognate antigen. PVI.V6-14 binds a novel, conserved epitope on the N-terminal domain (NTD) and neutralizes the original viral strain. PVI.V6-14 also binds the newly emerged variants B.1.1.7 (alpha), B.1.351 (beta), B.1.617.2 (delta), and B.1.1.529 (omicron). Given that this antibody was likely already present in the donor prior to vaccination, we believe that this antibody class could potentially "keep up" with the new variants, should they continue to emerge, by undergoing somatic hypermutation and affinity maturation.
Keywords: adaptive immunity; coronavirus; electron microscopy; monoclonal antibodies; vaccine.
Conflict of interest statement
The authors declare a conflict of interest. The Icahn School of Medicine at Mount Sinai has filed patent applications relating to SARS-CoV-2 serological assays and NDV-based SARS-CoV-2 vaccines which list Florian Krammer as co-inventor. Viviana Simon and Fatima Amanat are also listed on the serological assay patent application as co-inventors. Mount Sinai has spun out a company, Kantaro, to market serological tests for SARS-CoV-2. Florian Krammer has consulted for Merck and Pfizer (before 2020), and is currently consulting for Pfizer, Seqirus and Avimex. The Krammer laboratory is also collaborating with Pfizer on animal models of SARS-CoV-2. Ali Ellebedy has consulted for InBios and Fimbrion Therapeutics (before 2021) and is currently a consultant for Mubadala Investment Company. The Ellebedy laboratory received funding under sponsored research agreements that are unrelated to the data presented in the current study from Emergent BioSolutions and from AbbVie.
Figures


References
-
- Baden LR, El Sahly HM, Essink B, Kotloff K, Frey S, Novak R, Diemert D, Spector SA, Rouphael N, Creech CB, McGettigan J, Khetan S, Segall N, Solis J, Brosz A, Fierro C, Schwartz H, Neuzil K, Corey L, Gilbert P, Janes H, Follmann D, Marovich M, Mascola J, Polakowski L, Ledgerwood J, Graham BS, Bennett H, Pajon R, Knightly C, Leav B, Deng W, Zhou H, Han S, Ivarsson M, Miller J, Zaks T, COVE Study Group. 2021. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N Engl J Med 384:403–416. doi: 10.1056/NEJMoa2035389. - DOI - PMC - PubMed
-
- Polack FP, Thomas SJ, Kitchin N, Absalon J, Gurtman A, Lockhart S, Perez JL, Pérez Marc G, Moreira ED, Zerbini C, Bailey R, Swanson KA, Roychoudhury S, Koury K, Li P, Kalina WV, Cooper D, Frenck RW, Jr, Hammitt LL, Türeci Ö, Nell H, Schaefer A, Ünal S, Tresnan DB, Mather S, Dormitzer PR, Şahin U, Jansen KU, Gruber WC, C4591001 Clinical Trial Group. 2020. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N Engl J Med 383:2603–2615. doi: 10.1056/NEJMoa2034577. - DOI - PMC - PubMed
-
- Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu N-H, Nitsche A, Müller MA, Drosten C, Pöhlmann S. 2020. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271–280.e8. doi: 10.1016/j.cell.2020.02.052. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

