Chicken liver is a potential reservoir of bacteriophages and phage-derived particles containing antibiotic resistance genes
- PMID: 35485188
- PMCID: PMC9437878
- DOI: 10.1111/1751-7915.14056
Chicken liver is a potential reservoir of bacteriophages and phage-derived particles containing antibiotic resistance genes
Abstract
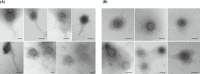
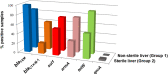
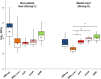
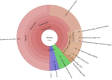
Poultry meat production is one of the most important agri-food industries in the world. The selective pressure exerted by widespread prophylactic or therapeutic use of antibiotics in intensive chicken farming favours the development of drug resistance in bacterial populations. Chicken liver, closely connected with the intestinal tract, has been directly involved in food-borne infections and found to be contaminated with pathogenic bacteria, including Campylobacter and Salmonella. In this study, 74 chicken livers, divided into sterile and non-sterile groups, were analysed, not only for microbial indicators but also for the presence of phages and phage particles containing antibiotic resistance genes (ARGs). Both bacteria and phages were detected in liver tissues, including those dissected under sterile conditions. The phages were able to infect Escherichia coli and showed a Siphovirus morphology. The chicken livers contained from 103 to 106 phage particles per g, which carried a range of ARGs (blaTEM , blaCTx-M-1 , sul1, qnrA, armA and tetW) detected by qPCR. The presence of phages in chicken liver, mostly infecting E. coli, was confirmed by metagenomic analysis, although this technique was not sufficiently sensitive to identify ARGs. In addition, ARG-carrying phages were detected in chicken faeces by qPCR in a previous study of the group. Comparison of the viromes of faeces and liver showed a strong coincidence of species, which suggests that the phages found in the liver originate in faeces. These findings suggests that phages, like bacteria, can translocate from the gut to the liver, which may therefore constitute a potential reservoir of antibiotic resistance genes.
© 2022 The Authors. Microbial Biotechnology published by Society for Applied Microbiology and John Wiley & Sons Ltd.
Conflict of interest statement
None declared.
Figures

Similar articles
-
Infectious phage particles packaging antibiotic resistance genes found in meat products and chicken feces.Sci Rep. 2019 Sep 16;9(1):13281. doi: 10.1038/s41598-019-49898-0. Sci Rep. 2019. PMID: 31527758 Free PMC article.
-
Antibiotic resistance genes in phage particles isolated from human faeces and induced from clinical bacterial isolates.Int J Antimicrob Agents. 2018 Mar;51(3):434-442. doi: 10.1016/j.ijantimicag.2017.11.014. Epub 2017 Nov 24. Int J Antimicrob Agents. 2018. PMID: 29180282
-
CrAss-like phages are suitable indicators of antibiotic resistance genes found in abundance in fecally polluted samples.Environ Pollut. 2024 Oct 15;359:124713. doi: 10.1016/j.envpol.2024.124713. Epub 2024 Aug 10. Environ Pollut. 2024. PMID: 39134166
-
[Soil phage and their mediation on the horizontal transfer of antibiotic resistance genes: A review].Ying Yong Sheng Tai Xue Bao. 2021 Jun;32(6):2267-2274. doi: 10.13287/j.1001-9332.202106.031. Ying Yong Sheng Tai Xue Bao. 2021. PMID: 34212633 Review. Chinese.
-
Phage-antibiotic combinations: a promising approach to constrain resistance evolution in bacteria.Ann N Y Acad Sci. 2021 Jul;1496(1):23-34. doi: 10.1111/nyas.14533. Epub 2020 Nov 11. Ann N Y Acad Sci. 2021. PMID: 33175408 Review.
Cited by
-
The Microbiome and Metabolic Dysfunction-Associated Steatotic Liver Disease.Int J Mol Sci. 2025 Mar 22;26(7):2882. doi: 10.3390/ijms26072882. Int J Mol Sci. 2025. PMID: 40243472 Free PMC article. Review.
-
Experimental evolution at ecological scales allows linking of viral genotypes to specific host strains.ISME J. 2024 Jan 8;18(1):wrae208. doi: 10.1093/ismejo/wrae208. ISME J. 2024. PMID: 39579348 Free PMC article.
-
Non-Canonical Aspects of Antibiotics and Antibiotic Resistance.Antibiotics (Basel). 2024 Jun 17;13(6):565. doi: 10.3390/antibiotics13060565. Antibiotics (Basel). 2024. PMID: 38927231 Free PMC article. Review.
-
Bioinputs from Eugenia dysenterica DC. (Myrtaceae): Optimization of Ultrasound-Assisted Extraction and Assessment of Antioxidant, Antimicrobial, and Antibiofilm Activities.Molecules. 2025 Feb 28;30(5):1115. doi: 10.3390/molecules30051115. Molecules. 2025. PMID: 40076338 Free PMC article.
-
Prospects and Challenges of Bacteriophage Substitution for Antibiotics in Livestock and Poultry Production.Biology (Basel). 2024 Jan 4;13(1):28. doi: 10.3390/biology13010028. Biology (Basel). 2024. PMID: 38248459 Free PMC article. Review.
References
-
- Abdelhamid, M.K. , Quijada, N.M. , Dzieciol, M. , Hatfaludi, T. , Bilic, I. , Selberherr, E. , et al. (2020) Co‐infection of chicken layers with Histomonas meleagridis and avian pathogenic Escherichia coli is associated with dysbiosis, cecal colonization and translocation of the bacteria from the gut lumen. Front Microbiol 11: 586437. - PMC - PubMed
-
- Anonymous . (2000) ISO 10705‐2: Water quality. Detection and enumeration of bacteriophages ‐part 2: enumeration of somatic coliphages.
-
- Antimicrobial Resistance in the Environment Summary Report of an FAO Meeting of Experts FAO Antimicrobial Resistance Working Group (2018).
-
- Awad, W.A. , Dublecz, F. , Hess, C. , Dublecz, K. , Khayal, B. , Aschenbach, J.R. , and Hess, M. (2016) Campylobacter jejuni colonization promotes the translocation of Escherichia coli to extra‐intestinal organs and disturbs the short‐chain fatty acids profiles in the chicken gut. Poult Sci 95: 2259–2265. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

