Melanin biopolymer synthesis using a new melanogenic strain of Flavobacterium kingsejongi and a recombinant strain of Escherichia coli expressing 4-hydroxyphenylpyruvate dioxygenase from F. kingsejongi
- PMID: 35501871
- PMCID: PMC9063278
- DOI: 10.1186/s12934-022-01800-w
Melanin biopolymer synthesis using a new melanogenic strain of Flavobacterium kingsejongi and a recombinant strain of Escherichia coli expressing 4-hydroxyphenylpyruvate dioxygenase from F. kingsejongi
Abstract
Background: Melanins are a heterologous group of biopolymeric pigments synthesized by diverse prokaryotes and eukaryotes and are widely utilized as bioactive materials and functional polymers in the biotechnology industry. Here, we report the high-level melanin production using a new melanogenic Flavobacterium kingsejongi strain and a recombinant Escherichia coli overexpressing F. kingsejongi 4-hydroxyphenylpyruvate dioxygenase (HPPD).
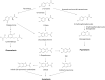
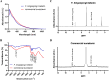
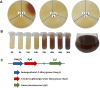
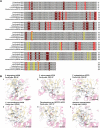
Results: Melanin synthesis of F. kingsejongi strain was confirmed via melanin synthesis inhibition test, melanin solubility test, genome analysis, and structural analysis of purified melanin from both wild-type F. kingsejongi and recombinant E. coli expressing F. kingsejongi HPPD. The activity of F. kingsejongi HPPD was demonstrated via in vitro assays with 6 × His-tagged and native forms of HPPD. The specific activity of F. kingsejongi HPPD was 1.2 ± 0.03 μmol homogentisate/min/mg-protein. Bioreactor fermentation of F. kingsejongi produced a large amount of melanin with a titer of 6.07 ± 0.32 g/L, a conversion yield of 60% (0.6 ± 0.03 g melanin per gram tyrosine), and a productivity of 0.03 g/L·h, indicating its potential for industrial melanin production. Additionally, bioreactor fermentation of recombinant E. coli expressing F. kingsejongi HPPD produced melanin at a titer of 3.76 ± 0.30 g/L, a conversion yield of 38% (0.38 ± 0.03 g melanin per gram tyrosine), and a productivity of 0.04 g/L·h.
Conclusions: Both strains showed sufficiently high fermentation capability to indicate their potential as platform strains for large-scale bacterial melanin production. Furthermore, F. kingsejongi strain could serve as a model to elucidate the regulation of melanin biosynthesis pathway and its networks with other cellular pathways, and to understand the cellular responses of melanin-producing bacteria to environmental changes, including nutrient starvation and other stresses.
Keywords: 4-Hydroxyphenylpyruvate dioxygenase; Flavobacterium kingsejongi; Melanin.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

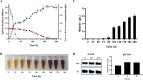

References
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

