Why Plasmodium vivax and Plasmodium falciparum are so different? A tale of two clades and their species diversities
- PMID: 35505356
- PMCID: PMC9066883
- DOI: 10.1186/s12936-022-04130-9
Why Plasmodium vivax and Plasmodium falciparum are so different? A tale of two clades and their species diversities
Abstract
The global malaria burden sometimes obscures that the genus Plasmodium comprises diverse clades with lineages that independently gave origin to the extant human parasites. Indeed, the differences between the human malaria parasites were highlighted in the classical taxonomy by dividing them into two subgenera, the subgenus Plasmodium, which included all the human parasites but Plasmodium falciparum that was placed in its separate subgenus, Laverania. Here, the evolution of Plasmodium in primates will be discussed in terms of their species diversity and some of their distinct phenotypes, putative molecular adaptations, and host-parasite biocenosis. Thus, in addition to a current phylogeny using genome-level data, some specific molecular features will be discussed as examples of how these parasites have diverged. The two subgenera of malaria parasites found in primates, Plasmodium and Laverania, reflect extant monophyletic groups that originated in Africa. However, the subgenus Plasmodium involves species in Southeast Asia that were likely the result of adaptive radiation. Such events led to the Plasmodium vivax lineage. Although the Laverania species, including P. falciparum, has been considered to share "avian characteristics," molecular traits that were likely in the common ancestor of primate and avian parasites are sometimes kept in the Plasmodium subgenus while being lost in Laverania. Assessing how molecular traits in the primate malaria clades originated is a fundamental science problem that will likely provide new targets for interventions. However, given that the genus Plasmodium is paraphyletic (some descendant groups are in other genera), understanding the evolution of malaria parasites will benefit from studying "non-Plasmodium" Haemosporida.
Keywords: Chitinases; Laverania; Malaria; Merozoite surface protein; Molecular clock; PfRH5-PfCyRPA-PfRipr complex; Phylogenomics; Plasmodium phylogeny.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

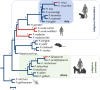



References
-
- Coatney GR, Collins WE, Warren M, Contacos PG. The primate malarias. 1. Bethesda: US National Institute of Allergy and Infectious Diseases; 1971.
-
- Garnham PCC. Malaria parasites and other Haemosporidia. Oxford: Blackwell Science Ltd; 1966.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

