A murine periodontitis model using coaggregation between human pathogens and a predominant mouse oral commensal bacterium
- PMID: 35505575
- PMCID: PMC9064776
- DOI: 10.5051/jpis.2104360218
A murine periodontitis model using coaggregation between human pathogens and a predominant mouse oral commensal bacterium
Abstract
Purpose: C57BL/6 mice, which are among the most common backgrounds for genetically engineered mice, are resistant to the induction of periodontitis by oral infection with periodontal pathogens. This study aimed to develop a periodontitis model in C57BL/6 mice using coaggregation between human pathogens and the mouse oral commensal Streptococcus danieliae (Sd).
Methods: The abilities of Porphyromonas gingivalis ATCC 33277 (Pg33277), P. gingivalis ATCC 49417 (Pg49417), P. gingivalis KUMC-P4 (PgP4), Fusobacterium nucleatum subsp. nucleatum ATCC 25586 (Fnn), and F. nucleatum subsp. animalis KCOM 1280 (Fna) to coaggregate with Sd were tested by a sedimentation assay. The Sd-noncoaggregating Pg33277 and 2 Sd-coaggregating strains, PgP4 and Fna, were chosen for animal experiments. Eighty C57BL/6 mice received oral gavage with Sd once and subsequently received vehicle alone (sham), Fna, Pg33277, PgP4, or Fna+PgP4 6 times at 2-day intervals. Mice were evaluated at 5 or 8 weeks after the first gavage of human strains.
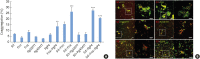
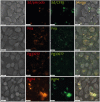
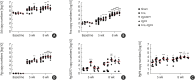
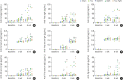
Results: Fnn, Fna, and PgP4 efficiently coaggregated with Sd, but Pg33277 and Pg49417 did not. Alveolar bone loss was significantly higher in the PgP4 group at both time points (weeks 5 and 8) and in all experimental groups at week 8 compared with the sham group. The PgP4 group presented greater alveolar bone loss than the other experimental groups at both time points. A higher degree of alveolar bone loss accompanied higher bacterial loads in the oral cavity, the invasion of not only PgP4 but also Sd and Fna, and the serum antibody responses to these bacteria.
Conclusions: Periodontitis was successfully induced in C57BL/6 mice by oral infection with a P. gingivalis strain that persists in the oral cavity through coaggregation with a mouse oral commensal bacterium. This new model will be useful for studying the role of human oral bacteria-host interactions in periodontitis using genetically engineered mice.
Keywords:
Animal models; Mice; Periodontitis;
Copyright © 2022. Korean Academy of Periodontology.
Conflict of interest statement
No potential conflict of interest relevant to this article was reported.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

