miR-6077 promotes cisplatin/pemetrexed resistance in lung adenocarcinoma via CDKN1A/cell cycle arrest and KEAP1/ferroptosis pathways
- PMID: 35505963
- PMCID: PMC9035384
- DOI: 10.1016/j.omtn.2022.03.020
miR-6077 promotes cisplatin/pemetrexed resistance in lung adenocarcinoma via CDKN1A/cell cycle arrest and KEAP1/ferroptosis pathways
Abstract
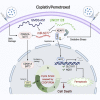
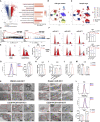
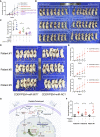
Lung adenocarcinoma (LUAD) is one of the most common malignancies worldwide. Combination chemotherapy with cisplatin (CDDP) plus pemetrexed (PEM) remains the predominant therapeutic regimen; however, chemoresistance greatly limits its curative potential. Here, through CRISPR-Cas9 screening, we identified miR-6077 as a key driver of CDDP/PEM resistance in LUAD. Functional experiments verified that ectopic overexpression of miR-6077 desensitized LUAD cells to CDDP/PEM in both cell lines and patient-derived xenograft models. Through RNA sequencing in cells and single-cell sequencing of samples from patients with CDDP/PEM treatments, we observed CDDP/PEM-induced upregulation of CDKN1A and KEAP1, which in turn activated cell-cycle arrest and ferroptosis, respectively, thus leading to cell death. Through miRNA pull-down, we identified and validated that miR-6077 targets CDKN1A and KEAP1. Furthermore, we demonstrated that miR-6077 protects LUAD cells from cell death induced by CDDP/PEM via CDKN1A-CDK1-mediated cell-cycle arrest and KEAP1-NRF2-SLC7A11/NQO1-mediated ferroptosis, thus resulting in chemoresistance in multiple LUAD cells both in vitro and in vivo. Moreover, we found that GMDS-AS1 and LINC01128 sensitized LUAD cells to CDDP/PEM by sponging miR-6077. Collectively, these results imply the critical role of miR-6077 in LUAD's sensitivity to CDDP/PEM, thus providing a novel therapeutic strategy for overcoming chemoresistance in clinical practice.
Keywords: CRISPR-Cas9; MT: Non-coding RNAs; RNA-seq; cell-cycle arrest; chemoresistance; cisplatin; ferroptosis; homology-directed repair; lncRNA; miRNA; pemetrexed.
© 2022 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Circular RNA CDR1-AS contributes to pemetrexed and cisplatin chemoresistance through EGFR/PI3K signaling pathway in lung adenocarcinoma.Biomed Pharmacother. 2020 Mar;123:109771. doi: 10.1016/j.biopha.2019.109771. Epub 2019 Dec 24. Biomed Pharmacother. 2020. PMID: 31881486
-
CircP4HB regulates ferroptosis via SLC7A11-mediated glutathione synthesis in lung adenocarcinoma.Transl Lung Cancer Res. 2022 Mar;11(3):366-380. doi: 10.21037/tlcr-22-138. Transl Lung Cancer Res. 2022. PMID: 35399564 Free PMC article.
-
Randomized phase II study of chemoradiotherapy with cisplatin + S-1 versus cisplatin + pemetrexed for locally advanced non-squamous non-small cell lung cancer: SPECTRA study.Lung Cancer. 2020 Mar;141:64-71. doi: 10.1016/j.lungcan.2020.01.008. Epub 2020 Jan 10. Lung Cancer. 2020. PMID: 31955002 Clinical Trial.
-
miRNA-378 reverses chemoresistance to cisplatin in lung adenocarcinoma cells by targeting secreted clusterin.Sci Rep. 2016 Jan 19;6:19455. doi: 10.1038/srep19455. Sci Rep. 2016. PMID: 26781643 Free PMC article.
-
Exosomal microRNA-506 inhibits biological activity of lung adenocarcinoma cells and increases sensitivity to cisplatin-based hyperthermia.Cell Signal. 2022 Dec;100:110469. doi: 10.1016/j.cellsig.2022.110469. Epub 2022 Sep 14. Cell Signal. 2022. PMID: 36115547
Cited by
-
STK11 (LKB1) mutation suppresses ferroptosis in lung adenocarcinoma by facilitating monounsaturated fatty acid synthesis.Open Med (Wars). 2024 Jan 6;19(1):20230845. doi: 10.1515/med-2023-0845. eCollection 2024. Open Med (Wars). 2024. PMID: 38205151 Free PMC article.
-
Regulation of Ferroptosis in Lung Adenocarcinoma.Int J Mol Sci. 2023 Sep 27;24(19):14614. doi: 10.3390/ijms241914614. Int J Mol Sci. 2023. PMID: 37834062 Free PMC article. Review.
-
MNT inhibits lung adenocarcinoma ferroptosis and chemosensitivity by suppressing SAT1.Commun Biol. 2024 Jun 3;7(1):680. doi: 10.1038/s42003-024-06373-5. Commun Biol. 2024. PMID: 38831092 Free PMC article.
-
Recent advances in engineered exosome-based therapies for ocular vascular disease.J Nanobiotechnology. 2025 Jul 19;23(1):526. doi: 10.1186/s12951-025-03589-3. J Nanobiotechnology. 2025. PMID: 40684186 Free PMC article. Review.
-
Hypoxia Modulates Radiosensitivity and Response to Different Radiation Qualities in A549 Non-Small Cell Lung Cancer (NSCLC) Cells.Int J Mol Sci. 2024 Jan 13;25(2):1010. doi: 10.3390/ijms25021010. Int J Mol Sci. 2024. PMID: 38256084 Free PMC article.
References
-
- Sung H., Ferlay J., Siegel R.L., Laversanne M., Soerjomataram I., Jemal A., Bray F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021;71:209–249. - PubMed
-
- Arbour K.C., Riely G.J. Systemic therapy for locally advanced and metastatic non-small cell lung cancer: a review. JAMA. 2019;322:764–774. - PubMed
-
- Fennell D.A., Summers Y., Cadranel J., Benepal T., Christoph D.C., Lal R., Das M., Maxwell F., Visseren-Grul C., Ferry D. Cisplatin in the modern era: the backbone of first-line chemotherapy for non-small cell lung cancer. Cancer Treat. Rev. 2016;44:42–50. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

