CORP: Gene delivery into murine skeletal muscle using in vivo electroporation
- PMID: 35511722
- PMCID: PMC9236869
- DOI: 10.1152/japplphysiol.00088.2022
CORP: Gene delivery into murine skeletal muscle using in vivo electroporation
Abstract
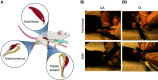
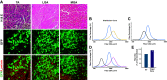
The strategy of gene delivery into skeletal muscles has provided exciting avenues in identifying new potential therapeutics toward muscular disorders and addressing basic research questions in muscle physiology through overexpression and knockdown studies. In vivo electroporation methodology offers a simple, rapidly effective technique for the delivery of plasmid DNA into postmitotic skeletal muscle fibers and the ability to easily explore the molecular mechanisms of skeletal muscle plasticity. The purpose of this review is to describe how to robustly electroporate plasmid DNA into different hindlimb muscles of rodent models. Furthermore, key parameters (e.g., voltage, hyaluronidase, and plasmid concentration) that contribute to the successful introduction of plasmid DNA into skeletal muscle fibers will be discussed. In addition, details on processing tissue for immunohistochemistry and fiber cross-sectional area (CSA) analysis will be outlined. The overall goal of this review is to provide the basic and necessary information needed for successful implementation of in vivo electroporation of plasmid DNA and thus open new avenues of discovery research in skeletal muscle physiology.
Keywords: atrophy; electroporation; gene transfer; hypertrophy; protein synthesis.
Conflict of interest statement
No conflicts of interest, financial or otherwise, are declared by the authors.
Figures



Similar articles
-
Electroporation of Plasmid DNA into Mouse Skeletal Muscle.J Vis Exp. 2022 Apr 6;(182). doi: 10.3791/63916. J Vis Exp. 2022. PMID: 35467646
-
A Critical Review of Electroporation as A Plasmid Delivery System in Mouse Skeletal Muscle.Int J Mol Sci. 2019 Jun 6;20(11):2776. doi: 10.3390/ijms20112776. Int J Mol Sci. 2019. PMID: 31174257 Free PMC article. Review.
-
Optimizing plasmid-based gene transfer for investigating skeletal muscle structure and function.Mol Ther. 2006 Apr;13(4):795-803. doi: 10.1016/j.ymthe.2005.09.019. Epub 2005 Nov 23. Mol Ther. 2006. PMID: 16309967
-
A comprehensive study of optimal conditions for naked plasmid DNA transfer into skeletal muscle by electroporation.J Gene Med. 2005 Sep;7(9):1235-45. doi: 10.1002/jgm.765. J Gene Med. 2005. PMID: 15880605
-
Plasmid DNA gene therapy by electroporation: principles and recent advances.Curr Gene Ther. 2011 Dec;11(6):447-56. doi: 10.2174/156652311798192860. Curr Gene Ther. 2011. PMID: 22023474 Review.
Cited by
-
Skeletal muscle contraction kinetics and AMPK responses are modulated by the adenine nucleotide degrading enzyme AMPD1.J Appl Physiol (1985). 2022 Nov 1;133(5):1055-1066. doi: 10.1152/japplphysiol.00035.2022. Epub 2022 Sep 15. J Appl Physiol (1985). 2022. PMID: 36107988 Free PMC article.
-
Reprogramming of cis-regulatory networks during skeletal muscle atrophy in male mice.Nat Commun. 2023 Oct 18;14(1):6581. doi: 10.1038/s41467-023-42313-3. Nat Commun. 2023. PMID: 37853001 Free PMC article.
-
Combining In Vivo 2-Photon Imaging with Photoactivatable Fluorescent Labeling Shows Low Rates of Mitochondrial Dynamics in Skeletal Muscle.Med Sci Sports Exerc. 2025 May 1:10.1249/MSS.0000000000003748. doi: 10.1249/MSS.0000000000003748. Online ahead of print. Med Sci Sports Exerc. 2025. PMID: 40310022 Free PMC article.
-
Response of UBR-box E3 ubiquitin ligases and protein quality control pathways to perturbations in protein synthesis and skeletal muscle size.bioRxiv [Preprint]. 2025 Jul 27:2025.07.23.666188. doi: 10.1101/2025.07.23.666188. bioRxiv. 2025. PMID: 40777329 Free PMC article. Preprint.
-
Mechanisms of mechanical overload-induced skeletal muscle hypertrophy: current understanding and future directions.Physiol Rev. 2023 Oct 1;103(4):2679-2757. doi: 10.1152/physrev.00039.2022. Epub 2023 Jun 29. Physiol Rev. 2023. PMID: 37382939 Free PMC article. Review.
References
-
- Bodine SC, Stitt TN, Gonzalez M, Kline WO, Stover GL, Bauerlein R, Zlotchenko E, Scrimgeour A, Lawrence JC, Glass DJ, Yancopoulos GD. Akt/mTOR pathway is a crucial regulator of skeletal muscle hypertrophy and can prevent muscle atrophy in vivo. Nat Cell Biol 3: 1014–1019, 2001. doi:10.1038/ncb1101-1014. - DOI - PubMed
-
- Bodine SC, Latres E, Baumhueter S, Lai VK, Nunez L, Clarke BA, Poueymirou WT, Panaro FJ, Na E, Dharmarajan K, Pan ZQ, Valenzuela DM, DeChiara TM, Stitt TN, Yancopoulos GD, Glass DJ. Identification of ubiquitin ligases required for skeletal muscle atrophy. Science 294: 1704–1708, 2001. doi:10.1126/science.1065874. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

