Circadian entrainment in Arabidopsis
- PMID: 35512209
- PMCID: PMC9516740
- DOI: 10.1093/plphys/kiac204
Circadian entrainment in Arabidopsis
Abstract
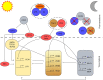
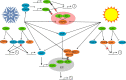
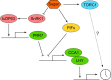
Circadian clocks coordinate physiology and development as an adaption to the oscillating day/night cycle caused by the rotation of Earth on its axis and the changing length of day and night away from the equator caused by orbiting the sun. Circadian clocks confer advantages by entraining to rhythmic environmental cycles to ensure that internal events within the plant occur at the correct time with respect to the cyclic external environment. Advances in determining the structure of circadian oscillators and the pathways that allow them to respond to light, temperature, and metabolic signals have begun to provide a mechanistic insight to the process of entrainment in Arabidopsis (Arabidopsis thaliana). We describe the concepts of entrainment and how it occurs. It is likely that a thorough mechanistic understanding of the genetic and physiological basis of circadian entrainment will provide opportunities for crop improvement.
© The Author(s) 2022. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Figures

Similar articles
-
BIG Regulates Dynamic Adjustment of Circadian Period in Arabidopsis thaliana.Plant Physiol. 2018 Sep;178(1):358-371. doi: 10.1104/pp.18.00571. Epub 2018 Jul 11. Plant Physiol. 2018. PMID: 29997180 Free PMC article.
-
From a repressilator-based circadian clock mechanism to an external coincidence model responsible for photoperiod and temperature control of plant architecture in Arabodopsis thaliana.Biosci Biotechnol Biochem. 2013;77(1):10-6. doi: 10.1271/bbb.120765. Epub 2013 Jan 7. Biosci Biotechnol Biochem. 2013. PMID: 23291766 Review.
-
Arabidopsis EARLY FLOWERING 3 controls temperature responsiveness of the circadian clock independently of the evening complex.J Exp Bot. 2022 Jan 27;73(3):1049-1061. doi: 10.1093/jxb/erab473. J Exp Bot. 2022. PMID: 34698833
-
Circadian Entrainment in Arabidopsis by the Sugar-Responsive Transcription Factor bZIP63.Curr Biol. 2018 Aug 20;28(16):2597-2606.e6. doi: 10.1016/j.cub.2018.05.092. Epub 2018 Aug 2. Curr Biol. 2018. PMID: 30078562 Free PMC article.
-
Illuminating the Arabidopsis circadian epigenome: Dynamics of histone acetylation and deacetylation.Curr Opin Plant Biol. 2022 Oct;69:102268. doi: 10.1016/j.pbi.2022.102268. Epub 2022 Jul 31. Curr Opin Plant Biol. 2022. PMID: 35921796 Review.
Cited by
-
Focus on circadian rhythms.Plant Physiol. 2022 Sep 28;190(2):921-923. doi: 10.1093/plphys/kiac353. Plant Physiol. 2022. PMID: 35900174 Free PMC article. No abstract available.
-
Single-Nucleus Transcriptomics Reveals How Cell Type Shapes the Circadian Transcriptome of the Arabidopsis Leaf.bioRxiv [Preprint]. 2025 Jul 16:2025.06.12.659411. doi: 10.1101/2025.06.12.659411. bioRxiv. 2025. PMID: 40667004 Free PMC article. Preprint.
-
Parameterized resetting model captures dose-dependent entrainment of the mouse circadian clock.Nat Commun. 2025 Feb 6;16(1):1421. doi: 10.1038/s41467-025-56792-z. Nat Commun. 2025. PMID: 39915501 Free PMC article.
-
HOS15-mediated turnover of PRR7 enhances freezing tolerance.New Phytol. 2024 Nov;244(3):798-810. doi: 10.1111/nph.20062. Epub 2024 Aug 19. New Phytol. 2024. PMID: 39155726
-
The circadian clock controls temporal and spatial patterns of floral development in sunflower.Elife. 2023 Jan 13;12:e80984. doi: 10.7554/eLife.80984. Elife. 2023. PMID: 36637156 Free PMC article.
References
-
- Anderson SJ, Kramer MC, Gosai SJ, Yu X, Vandivier LE, Nelson ADL, Anderson ZD, Beilstein MA, Fray RG, Lyons E, et al. (2018) N6-methyladenosine inhibits local ribonucleolytic cleavage to tabilize mRNAs in Arabidopsis. Cell Rep 25: 1146–1157 - PubMed
-
- Balcerowicz M, Mahjoub M, Nguyen D, Lan H, Stoeckle D, Conde S, Jaeger KE, Wigge PA, Ezer D (2021) An early-morning gene network controlled by phytochromes and cryptochromes regulates photomorphogenesis pathways in Arabidopsis. Mol Plant 14: 983–996 - PubMed
-
- Box MS, Huang BE, Domijan M, Jaeger KE, Khattak AK, Yoo SJ, Sedivy EL, Jones DM, Hearn TJ, Webb AAR, et al. (2015) ELF3 controls thermoresponsive growth in Arabidopsis. Curr Biol 25: 194–199 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

