Comparative optimization of combinatorial CRISPR screens
- PMID: 35513429
- PMCID: PMC9072436
- DOI: 10.1038/s41467-022-30196-9
Comparative optimization of combinatorial CRISPR screens
Abstract
Combinatorial CRISPR technologies have emerged as a transformative approach to systematically probe genetic interactions and dependencies of redundant gene pairs. However, the performance of different functional genomic tools for multiplexing sgRNAs vary widely. Here, we generate and benchmark ten distinct pooled combinatorial CRISPR libraries targeting paralog pairs to optimize digenic knockout screens. Libraries composed of dual Streptococcus pyogenes Cas9 (spCas9), orthogonal spCas9 and Staphylococcus aureus (saCas9), and enhanced Cas12a from Acidaminococcus were evaluated. We demonstrate a combination of alternative tracrRNA sequences from spCas9 consistently show superior effect size and positional balance between the sgRNAs as a robust combinatorial approach to profile genetic interactions of multiple genes.
© 2022. The Author(s).
Conflict of interest statement
W.R.S. during the conduct of this research was or is a Board or SAB member and equity holder in Peloton Therapeutics, Ideaya Biosciences, Civetta Therapeutics, Scorpion Therapeutics, and Bluebird and has consulted for Array, Astex, Dynamo Therapeutics, Ipsen, PearlRiver Bio, Sanofi, and Servier; and receives research funding from Pfizer Pharmaceuticals, Merck, Ideaya Biosciences, Boehringer-Ingelheim, and Deerfield Management. T.I. is an equity holder of Scorpion Therapeutics and Odyssey Therapeutics. C.R.V. has received consulting fees from Flare Therapeutics, Roivant Sciences, and C4 Therapeutics, has served on the scientific advisory board of KSQ Therapeutics, Syros Pharmaceuticals, and Treeline Biosciences, has received research funding from Boehringer-Ingelheim and Treeline Biosciences and owns a stock option from Treeline Biosciences. The remaining authors declare no competing interests.
Figures




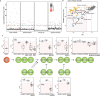
References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

