P-stereocontrolled synthesis of oligo(nucleoside N3'→O5' phosphoramidothioate)s - opportunities and limitations
- PMID: 35515667
- PMCID: PMC9056831
- DOI: 10.1039/d0ra04987e
P-stereocontrolled synthesis of oligo(nucleoside N3'→O5' phosphoramidothioate)s - opportunities and limitations
Abstract
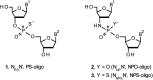
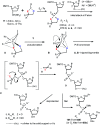
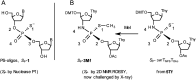
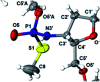
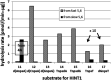
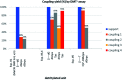
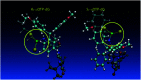
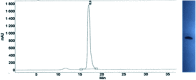
3'-N-(2-Thio-1,3,2-oxathiaphospholane) derivatives of 5'-O-DMT-3'-amino-2',3'-dideoxy-ribonucleosides (NOTP-N), that bear a 4,4-unsubstituted, 4,4-dimethyl, or 4,4-pentamethylene substituted oxathiaphospholane ring, were synthesized. Within these three series, NOTP-N differed by canonical nucleobases (i.e., AdeBz, CytBz, GuaiBu, or Thy). The monomers were chromatographically separated into P-diastereomers, which were further used to prepare NNPSN' dinucleotides (3), as well as short P-stereodefined oligo(deoxyribonucleoside N3'→O5' phosphoramidothioate)s (NPS-) and chimeric NPS/PO- and NPS/PS-oligomers. The condensation reaction for NOTP-N monomers was found to be 5-6 times slower than the analogous OTP derivatives. When the 5'-end nucleoside of a growing oligomer adopts a C3'-endo conformation, a conformational 'clash' with the incoming NOTP-N monomer takes place, which is a main factor decreasing the repetitive yield of chain elongation. Although both isomers of NNPSN' were digested by the HINT1 phosphoramidase enzyme, the isomers hydrolyzed at a faster rate were tentatively assigned the R P absolute configuration. This assignment is supported by X-ray analysis of the protected dinucleotide DMTdGiBu NPSMeTOAc, which is P-stereoequivalent to the hydrolyzed faster P-diastereomer of dGNPST.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures
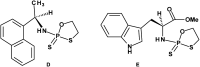

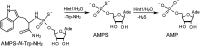


Similar articles
-
Synthesis and hybridizing properties of P-stereodefined chimeric [PS]-{DNA:RNA} and [PS]-{DNA:(2'-OMe)-RNA} oligomers.RSC Adv. 2022 Sep 21;12(41):26815-26824. doi: 10.1039/d2ra04855h. eCollection 2022 Sep 16. RSC Adv. 2022. PMID: 36320848 Free PMC article.
-
P-Stereodefined phosphorothioate analogs of glycol nucleic acids-synthesis and structural properties.RSC Adv. 2018 Jul 11;8(44):24942-24952. doi: 10.1039/c8ra05568h. eCollection 2018 Jul 9. RSC Adv. 2018. PMID: 35542141 Free PMC article.
-
Thermal stability and conformation of antiparallel duplexes formed by P-stereodefined phosphorothioate DNA/LNA chimeric oligomers with DNA and RNA matrices.Org Biomol Chem. 2015 Oct 21;13(39):10032-40. doi: 10.1039/c5ob01474c. Org Biomol Chem. 2015. PMID: 26293357
-
Synthesis of phosphorothioate oligonucleotides with stereodefined phosphorothioate linkages.Curr Protoc Nucleic Acid Chem. 2003 Oct;Chapter 4:Unit 4.17. doi: 10.1002/0471142700.nc0417s14. Curr Protoc Nucleic Acid Chem. 2003. PMID: 18428907 Review.
-
Oligonucleotide n3'-->p5' phosphoramidates and thio-phoshoramidates as potential therapeutic agents.Chem Biodivers. 2010 Mar;7(3):477-93. doi: 10.1002/cbdv.200900187. Chem Biodivers. 2010. PMID: 20232321 Review.
Cited by
-
Synthesis and hybridizing properties of P-stereodefined chimeric [PS]-{DNA:RNA} and [PS]-{DNA:(2'-OMe)-RNA} oligomers.RSC Adv. 2022 Sep 21;12(41):26815-26824. doi: 10.1039/d2ra04855h. eCollection 2022 Sep 16. RSC Adv. 2022. PMID: 36320848 Free PMC article.
References
-
- Zamecnik P. C. Stephenson M. L. Proc. Natl. Acad. Sci. U. S. A. 1978;75:280. doi: 10.1073/pnas.75.1.280. - DOI - PMC - PubMed
- Kurreck J. Eur. J. Biochem. 2003;270:1628. doi: 10.1073/pnas.75.1.280. - DOI - PubMed
- Kole R. Krainer A. R. Altman S. Nat. Rev. Drug Discovery. 2012,;11:12. doi: 10.1073/pnas.75.1.280. - DOI - PMC - PubMed
-
- Eckstein F. Antisense Nucleic Acid Drug Dev. 2000;10:117. doi: 10.1089/oli.1.2000.10.117. - DOI - PubMed
- Guga P. Koziołkiewicz M. Chem. Biodiversity. 2011;8:1642. doi: 10.1002/cbdv.201100130. - DOI - PubMed
- Eckstein F. Nucleic Acid Ther. 2014;24:374. doi: 10.1089/nat.2014.0506. - DOI - PubMed
- Volk D. E. Lokesh G. L. R. Biomedicines. 2017;5:41. doi: 10.3390/biomedicines5030041. - DOI - PMC - PubMed
- Shen X. Corey D. R. Nucleic Acids Res. 2018;46:1584. doi: 10.1093/nar/gkx1239. - DOI - PMC - PubMed
-
- Stec W. J. Karwowski B. Boczkowska M. Guga P. Koziołkiewicz M. Sochacki M. Wieczorek M. Błaszczyk J. J. Am. Chem. Soc. 1998;120:7156. doi: 10.1021/ja973801j. - DOI
- Guga P. and Stec W. J., in Current Protocols in Nucleic Acid Chemistry, ed. S. L. Beaucage, D. E. Bergstrom, G. D. Glick and R. A. Jones, John Wiley and Sons, Hoboken, N.J., 2003, p. 4.17.1 - PubMed
-
- Wilk A. Grajkowski A. Phillips L. R. Beaucage S. L. J. Am. Chem. Soc. 2000;122:2149. doi: 10.1021/ja991773u. - DOI
- Oka N. Yamamoto M. Sato T. Wada T. J. Am. Chem. Soc. 2008;130:16031. doi: 10.1021/ja991773u. - DOI - PubMed
- Knouse K. W. deGruyter J. N. Schmidt M. A. Zheng B. Vantourout J. C. Kingston C. Mercer S. E. Mcdonald I. M. Olson R. E. Zhu Y. Hang C. Zhu J. Yuan C. Wang Q. Park P. Eastgate M. D. Baran P. S. Science. 2018;361:1234. doi: 10.1021/ja991773u. - DOI - PMC - PubMed
- Iwamoto N. Butler D. C. D. Svrzikapa N. Mohapatra S. Zlatev I. Sah D. W. Y. Meena Standley S. M. Lu G. Apponi L. H. Frank-Kamenetsky M. Zhang J. J. Vargeese C. Verdine G. L. Nat. Biotechnol. 2017;35:845. doi: 10.1021/ja991773u. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

