Application of the Adverse Outcome Pathway Concept to In Vitro Nephrotoxicity Assessment: Kidney Injury due to Receptor-Mediated Endocytosis and Lysosomal Overload as a Case Study
- PMID: 35516525
- PMCID: PMC9061999
- DOI: 10.3389/ftox.2022.864441
Application of the Adverse Outcome Pathway Concept to In Vitro Nephrotoxicity Assessment: Kidney Injury due to Receptor-Mediated Endocytosis and Lysosomal Overload as a Case Study
Abstract
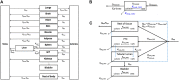
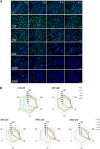
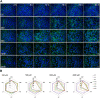
Application of adverse outcome pathways (AOP) and integration of quantitative in vitro to in vivo extrapolation (QIVIVE) may support the paradigm shift in toxicity testing to move from apical endpoints in test animals to more mechanism-based in vitro assays. Here, we developed an AOP of proximal tubule injury linking a molecular initiating event (MIE) to a cascade of key events (KEs) leading to lysosomal overload and ultimately to cell death. This AOP was used as a case study to adopt the AOP concept for systemic toxicity testing and risk assessment based on in vitro data. In this AOP, nephrotoxicity is thought to result from receptor-mediated endocytosis (MIE) of the chemical stressor, disturbance of lysosomal function (KE1), and lysosomal disruption (KE2) associated with release of reactive oxygen species and cytotoxic lysosomal enzymes that induce cell death (KE3). Based on this mechanistic framework, in vitro readouts reflecting each KE were identified. Utilizing polymyxin antibiotics as chemical stressors for this AOP, the dose-response for each in vitro endpoint was recorded in proximal tubule cells from rat (NRK-52E) and human (RPTEC/TERT1) in order to (1) experimentally support the sequence of key events (KEs), to (2) establish quantitative relationships between KEs as a basis for prediction of downstream KEs based on in vitro data reflecting early KEs and to (3) derive suitable in vitro points of departure for human risk assessment. Time-resolved analysis was used to support the temporal sequence of events within this AOP. Quantitative response-response relationships between KEs established from in vitro data on polymyxin B were successfully used to predict in vitro toxicity of other polymyxin derivatives. Finally, a physiologically based kinetic (PBK) model was utilized to transform in vitro effect concentrations to a human equivalent dose for polymyxin B. The predicted in vivo effective doses were in the range of therapeutic doses known to be associated with a risk for nephrotoxicity. Taken together, these data provide proof-of-concept for the feasibility of in vitro based risk assessment through integration of mechanistic endpoints and reverse toxicokinetic modelling.
Keywords: In vitro toxicity testing; QIVIVE; adverse outcome pathway (AOP); key event relationship; nephrotoxicity; risk assessment.
Copyright © 2022 Jarzina, Di Fiore, Ellinger, Reiser, Frank, Glaser, Wu, Taverne, Kramer and Mally.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
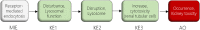




Similar articles
-
A case example of a radiation-relevant adverse outcome pathway to lung cancer.Int J Radiat Biol. 2021;97(1):68-84. doi: 10.1080/09553002.2019.1704913. Epub 2020 Jan 9. Int J Radiat Biol. 2021. PMID: 31846388
-
The adverse outcome pathway for rodent liver tumor promotion by sustained activation of the aryl hydrocarbon receptor.Regul Toxicol Pharmacol. 2015 Oct;73(1):172-90. doi: 10.1016/j.yrtph.2015.06.015. Epub 2015 Jul 2. Regul Toxicol Pharmacol. 2015. PMID: 26145830
-
Development of the Adverse Outcome Pathway (AOP): Chronic binding of antagonist to N-methyl-d-aspartate receptors (NMDARs) during brain development induces impairment of learning and memory abilities of children.Toxicol Appl Pharmacol. 2018 Sep 1;354:153-175. doi: 10.1016/j.taap.2018.02.024. Epub 2018 Mar 7. Toxicol Appl Pharmacol. 2018. PMID: 29524501 Free PMC article.
-
State of the art in non-animal approaches for skin sensitization testing: from individual test methods towards testing strategies.Arch Toxicol. 2016 Dec;90(12):2861-2883. doi: 10.1007/s00204-016-1842-4. Epub 2016 Sep 14. Arch Toxicol. 2016. PMID: 27629427 Review.
-
Building a Network of Adverse Outcome Pathways (AOPs) Incorporating the Tau-Driven AOP Toward Memory Loss (AOP429).J Alzheimers Dis Rep. 2022 Jun 7;6(1):271-296. doi: 10.3233/ADR-220015. eCollection 2022. J Alzheimers Dis Rep. 2022. PMID: 35891639 Free PMC article. Review.
Cited by
-
Development of an adverse outcome pathway network for nephrotoxicity.Arch Toxicol. 2024 Mar;98(3):929-942. doi: 10.1007/s00204-023-03637-7. Epub 2024 Jan 10. Arch Toxicol. 2024. PMID: 38197913 Free PMC article.
-
Oxide-Perovskites for Automotive Catalysts Biotransform and Induce Multicomponent Clearance and Hazard.ACS Nano. 2024 Nov 26;18(47):32672-32693. doi: 10.1021/acsnano.4c10135. Epub 2024 Nov 13. ACS Nano. 2024. PMID: 39537340
-
Mapping Adverse Outcome Pathways for Kidney Injury as a Basis for the Development of Mechanism-Based Animal-Sparing Approaches to Assessment of Nephrotoxicity.Front Toxicol. 2022 Jun 15;4:863643. doi: 10.3389/ftox.2022.863643. eCollection 2022. Front Toxicol. 2022. PMID: 35785263 Free PMC article.
-
Dextrin conjugation to colistin inhibits its toxicity, cellular uptake and acute kidney injury in vivo.RSC Pharm. 2024 Feb 5;1(1):68-79. doi: 10.1039/d3pm00014a. eCollection 2024 Apr 18. RSC Pharm. 2024. PMID: 38646595 Free PMC article.
References
LinkOut - more resources
Full Text Sources

