Investigating the biosynthesis of Sch-642305 in the fungus Phomopsis sp. CMU-LMA
- PMID: 35516937
- PMCID: PMC9055631
- DOI: 10.1039/d0ra05311b
Investigating the biosynthesis of Sch-642305 in the fungus Phomopsis sp. CMU-LMA
Abstract
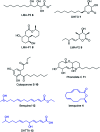
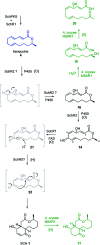
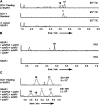
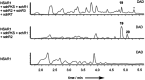
Sch-642305 is an unusual bicyclic 10-membered macrolide produced by the filamentous fungus Phomopsis sp. CMU-LMA for which no biosynthetic evidence exists. Here, we generate a draft genome sequence of the producing organism and discover the biosynthetic gene cluster responsible for formation of Sch-642305. Targeted gene disruptions together with reconstitution of the pathway in the heterologous host Aspergillus oryzae dissect key chemical steps and shed light on a series of oxidoreductions occuring in the pathway.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures


Similar articles
-
Isolation, structure elucidation and biological activity of metabolites from Sch-642305-producing endophytic fungus Phomopsis sp. CMU-LMA.Phytochemistry. 2011 Dec;72(18):2406-12. doi: 10.1016/j.phytochem.2011.08.010. Epub 2011 Sep 15. Phytochemistry. 2011. PMID: 21924749
-
Bioactive polyketides isolated from agar-supported fermentation of Phomopsis sp. CMU-LMA, taking advantage of the scale-up device, Platotex.Phytochemistry. 2013 Sep;93:170-5. doi: 10.1016/j.phytochem.2013.02.012. Epub 2013 Apr 9. Phytochemistry. 2013. PMID: 23578961
-
Biotransformation of natural compounds. Oxido-reduction of Sch-642305 by Aspergillus ochraceus ATCC 1009.Bioorg Med Chem Lett. 2011 Apr 15;21(8):2456-9. doi: 10.1016/j.bmcl.2011.02.063. Epub 2011 Feb 18. Bioorg Med Chem Lett. 2011. PMID: 21396813
-
Genome Editing Technology and Its Application Potentials in the Industrial Filamentous Fungus Aspergillus oryzae.J Fungi (Basel). 2021 Aug 5;7(8):638. doi: 10.3390/jof7080638. J Fungi (Basel). 2021. PMID: 34436177 Free PMC article. Review.
-
Strategies for the Enhancement of Secondary Metabolite Production via Biosynthesis Gene Cluster Regulation in Aspergillus oryzae.J Fungi (Basel). 2024 Apr 25;10(5):312. doi: 10.3390/jof10050312. J Fungi (Basel). 2024. PMID: 38786667 Free PMC article. Review.
References
-
- Mehta G. Shinde H. M. Tetrahedron Lett. 2005;46:6633–6636. doi: 10.1016/j.tetlet.2005.07.152. - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

