Conformational dynamics of superoxide dismutase (SOD1) in osmolytes: a molecular dynamics simulation study
- PMID: 35516947
- PMCID: PMC9055598
- DOI: 10.1039/d0ra02151b
Conformational dynamics of superoxide dismutase (SOD1) in osmolytes: a molecular dynamics simulation study
Abstract
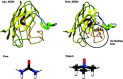
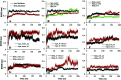
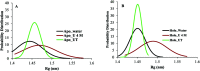
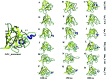
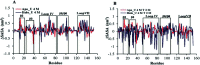
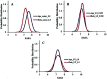
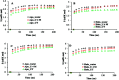
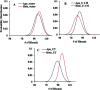
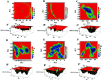
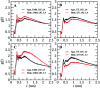
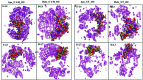
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disease caused by the misfolding of Cu, Zn superoxide dismutase (SOD1). Several earlier studies have shown that monomeric apo SOD1 undergoes significant local unfolding dynamics and is the predecessor for aggregation. Here, we have employed atomistic molecular dynamics (MD) simulations to study the structure and dynamics of monomeric apo and holo SOD1 in water, aqueous urea and aqueous urea-TMAO (trimethylamine oxide) solutions. Loop IV (zinc-binding loop) and loop VII (electrostatic loop) of holo SOD1 are considered as functionally important loops as they are responsible for the structural stability of holo SOD1. We found larger local unfolding of loop IV and VII of apo SOD1 as compared to holo SOD1 in water. Urea induced more unfolding in holo SOD1 than apo SOD1, whereas the stabilization of both the form of SOD1 was observed in ternary solution (i.e. water/urea/TMAO solution) but the extent of stabilization was higher in holo SOD1 than apo SOD1. The partially unfolded structures of apo SOD1 in water, urea and holo SOD1 in urea were identified by the exposure of the hydrophobic cores, which are highly dynamic and these may be the initial events of aggregation in SOD1. Our simulation studies support the formation of aggregates by means of the local unfolding of monomeric apo SOD1 as compared to holo SOD1 in water.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



Similar articles
-
TFE-induced local unfolding and fibrillation of SOD1: bridging the experiment and simulation studies.Biochem J. 2018 May 18;475(10):1701-1719. doi: 10.1042/BCJ20180085. Biochem J. 2018. PMID: 29686043
-
Solvent sensitivity of protein aggregation in Cu, Zn superoxide dismutase: a molecular dynamics simulation study.J Biomol Struct Dyn. 2018 Aug;36(10):2605-2617. doi: 10.1080/07391102.2017.1364670. Epub 2017 Aug 20. J Biomol Struct Dyn. 2018. PMID: 28782426
-
Insight Into Factors Influencing the Aggregation Process in Wild-Type and P66R Mutant SOD1: Computational and Spectroscopic Approaches.Proteins. 2025 Apr;93(4):885-907. doi: 10.1002/prot.26765. Epub 2024 Dec 6. Proteins. 2025. PMID: 39643934
-
The Role of Metal Binding in the Amyotrophic Lateral Sclerosis-Related Aggregation of Copper-Zinc Superoxide Dismutase.Molecules. 2017 Aug 29;22(9):1429. doi: 10.3390/molecules22091429. Molecules. 2017. PMID: 28850080 Free PMC article. Review.
-
Hydrogen Peroxide and Amyotrophic Lateral Sclerosis: From Biochemistry to Pathophysiology.Antioxidants (Basel). 2021 Dec 27;11(1):52. doi: 10.3390/antiox11010052. Antioxidants (Basel). 2021. PMID: 35052556 Free PMC article. Review.
Cited by
-
Flipping out: role of arginine in hydrophobic interactions and biological formulation design.Chem Sci. 2025 Mar 11;16(16):6780-6792. doi: 10.1039/d4sc08672d. eCollection 2025 Apr 16. Chem Sci. 2025. PMID: 40110519 Free PMC article.
-
Computational insight into in silico analysis and molecular dynamics simulation of the dimer interface residues of ALS-linked hSOD1 forms in apo/holo states: a combined experimental and bioinformatic perspective.3 Biotech. 2023 Mar;13(3):92. doi: 10.1007/s13205-023-03514-1. Epub 2023 Feb 21. 3 Biotech. 2023. PMID: 36845075 Free PMC article.
-
Amyloid fibrillation of the glaucoma associated myocilin protein is inhibited by epicatechin gallate (ECG).RSC Adv. 2022 Oct 14;12(45):29469-29481. doi: 10.1039/d2ra05061g. eCollection 2022 Oct 11. RSC Adv. 2022. PMID: 36320765 Free PMC article.
-
Structural insights into SOD1: from in silico and molecular dynamics to experimental analyses of ALS-associated E49K and R115G mutants.Front Mol Biosci. 2025 Feb 25;12:1532375. doi: 10.3389/fmolb.2025.1532375. eCollection 2025. Front Mol Biosci. 2025. PMID: 40070688 Free PMC article.
-
Copper toxicity and deficiency: the vicious cycle at the core of protein aggregation in ALS.Front Mol Neurosci. 2024 Jul 9;17:1408159. doi: 10.3389/fnmol.2024.1408159. eCollection 2024. Front Mol Neurosci. 2024. PMID: 39050823 Free PMC article. Review.
References
-
- Bruijn L. I. Houseweart M. K. Kato S. Anderson K. L. Anderson S. D. Ohama E. Reaume A. G. Scott R. W. Cleveland D. W. Aggregation and motor neuron toxicity of an ALS-linked SOD1 mutant independent from wild-type SOD1. Science. 1998;281(5384):1851–1854. doi: 10.1126/science.281.5384.1851. - DOI - PubMed
-
- Liu J. Lillo C. Jonsson P. A. Vande Velde C. Ward C. M. Miller T. M. Subramaniam J. R. Rothstein J. D. Marklund S. Andersen P. M. Brannstrom T. Gredal O. Wong P. C. Williams D. S. Cleveland D. W. Toxicity of familial ALS-linked SOD1 mutants from selective recruitment to spinal mitochondria. Neuron. 2004;43(1):5–17. doi: 10.1016/j.neuron.2004.06.016. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

