iTRAQ based proteomic analysis of PM2.5 induced lung damage
- PMID: 35517034
- PMCID: PMC9063425
- DOI: 10.1039/c9ra00252a
iTRAQ based proteomic analysis of PM2.5 induced lung damage
Abstract
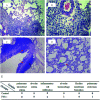
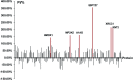
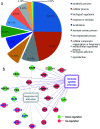
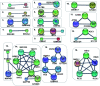
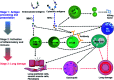
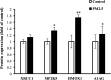
Haze pollution has become a global environmental problem, subsequently affecting air quality, climate, economy and human health. Notably, PM2.5 (particulate matter with an aerodynamic diameter less than 2.5 micrometers) significantly accounts for a variety of adverse health effects, in particular pulmonary diseases such as asthma and lung cancer. Clinical diagnosis and medical treatment of the lung damage caused by PM2.5 still remain significant challenges due to the lack of specific biomarkers and pathways. Here, we established a rat model of nonsurgical intratracheal instillation to investigate PM2.5 exposure and employed iTRAQ based analytical technique and bioinformatics tools to identify putative biomarkers and pathways. We identified 163 differentially expressed proteins (DEPs). Among these proteins, we screened six DEPs (HMOX1, MP2K5, XRCC1 E9PTZ7, KNT2 and A1AG) as the putative biomarkers, with significant differentially expressed levels (percentage increment > 140%). Pathway analysis indicated that calcium signaling, MAPK and PI3K/AKT might be involved in the process of PM2.5-induced lung damage. Western-blotting was used to verify DEPs in the AEC-II cell model for early diagnosis. In summary, our data can serve as fundamental research clues for further studies of PM2.5-induced toxicity in the lungs.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

References
-
- Huang R. J. Zhang Y. Bozzetti C. Ho K. F. Cao J. J. Han Y. Daellenbach K. R. Slowik J. G. Platt S. M. Canonaco F. Zotter P. Wolf R. Pieber S. M. Bruns E. A. Crippa M. Ciarelli G. Piazzalunga A. Schwikowski M. Abbaszade G. Schnelle-Kreis J. Zimmermann R. An Z. Szidat S. Baltensperger U. El Haddad I. Prevot A. S. Nature. 2014;514:218–222. doi: 10.1038/nature13774. - DOI - PubMed
LinkOut - more resources
Full Text Sources

