Inhibitory effects of UDP-glucuronosyltransferase (UGT) typical ligands against E. coli beta-glucuronidase (GUS)
- PMID: 35520305
- PMCID: PMC9054634
- DOI: 10.1039/d0ra02311f
Inhibitory effects of UDP-glucuronosyltransferase (UGT) typical ligands against E. coli beta-glucuronidase (GUS)
Abstract
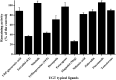
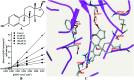
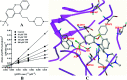
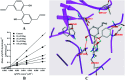
UDP-glucuronosyltransferases (UGTs) and β-glucuronidase (GUS) catalyze entirely distinct metabolism reactions. UGTs are responsible for the glucuronidation of a variety of drugs, endogenous and environmental chemicals, whereas GUS hydrolyzes glucuronides and liberates the parent substrates. Information on the overlap of ligand selectivity between UGT and GUS is essential for exploring the pharmacological or toxicological effects of the inhibitors of these two metabolic enzymes. This study is conducted to test whether UGTs and GUS share common ligands, by investigating the inhibitory effects towards E. coli GUS by a series of UGT typical substrates and inhibitors. Results showed that three typical ligands of UGTs, including two specific substrates (estradiol and trifluoperazine, E2 and TFP) and one selective inhibitor (magnolol, Mag), can inhibit the activity of GUS. Kinetic assays indicated that all the three UGT specific chemicals displayed competitive inhibition, with K i values of 31.4 (E2), 56.9 (TFP), and 16.6 μM (Mag). Docking studies further revealed that the three chemicals can enter the active sites of GUS by forming contacts with residues Glu-413, Trp-549, Asp-163, Tyr-472, Arg-562, or bound water. Our study indicates that ligand selectivity overlaps between UGTs and GUS, and some chemicals can act as co-inhibitors of these two metabolic enzymes. The pharmacological or toxicological effects of those co-inhibitors require further investigations.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures
Similar articles
-
Diverse effects of α-/β-estradiol on catalytic activities of human UDP-glucuronosyltransferases (UGT).J Steroid Biochem Mol Biol. 2023 Jan;225:106196. doi: 10.1016/j.jsbmb.2022.106196. Epub 2022 Sep 28. J Steroid Biochem Mol Biol. 2023. PMID: 36181991
-
Structure-inhibition relationship of podophyllotoxin (PT) analogues towards UDP-glucuronosyltransferase (UGT) isoforms.Pharmazie. 2015 Apr;70(4):239-43. Pharmazie. 2015. PMID: 26012253
-
A broad-spectrum substrate for the human UDP-glucuronosyltransferases and its use for investigating glucuronidation inhibitors.Int J Biol Macromol. 2021 Jun 1;180:252-261. doi: 10.1016/j.ijbiomac.2021.03.073. Epub 2021 Mar 16. Int J Biol Macromol. 2021. PMID: 33741369
-
A historical overview of the heterologous expression of mammalian UDP-glucuronosyltransferase isoforms over the past twenty years.Curr Drug Metab. 2005 Apr;6(2):141-60. doi: 10.2174/1389200053586127. Curr Drug Metab. 2005. PMID: 15853765 Review.
-
Structure and Protein-Protein Interactions of Human UDP-Glucuronosyltransferases.Front Pharmacol. 2016 Oct 24;7:388. doi: 10.3389/fphar.2016.00388. eCollection 2016. Front Pharmacol. 2016. PMID: 27822186 Free PMC article. Review.
Cited by
-
The Role of Gut Microbial β-Glucuronidase in Estrogen Reactivation and Breast Cancer.Front Cell Dev Biol. 2021 Aug 12;9:631552. doi: 10.3389/fcell.2021.631552. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34458248 Free PMC article. Review.
-
The tremendous clinical potential of the microbiota in the treatment of breast cancer: the next frontier.J Cancer Res Clin Oncol. 2023 Oct;149(13):12513-12534. doi: 10.1007/s00432-023-05014-4. Epub 2023 Jun 29. J Cancer Res Clin Oncol. 2023. PMID: 37382675 Free PMC article. Review.
-
In Vitro Ciclopirox Glucuronidation in Liver Microsomes from Humans and Various Experimental Animals.Eur J Drug Metab Pharmacokinet. 2024 Sep;49(5):619-629. doi: 10.1007/s13318-024-00907-2. Epub 2024 Jul 11. Eur J Drug Metab Pharmacokinet. 2024. PMID: 38990427
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

