H-lignin can be deposited independently of CINNAMYL ALCOHOL DEHYDROGENASE C and D in Arabidopsis
- PMID: 35522042
- PMCID: PMC9342963
- DOI: 10.1093/plphys/kiac210
H-lignin can be deposited independently of CINNAMYL ALCOHOL DEHYDROGENASE C and D in Arabidopsis
Abstract
Lignin contributes substantially to the recalcitrance of biomass toward saccharification. To circumvent this problem, researchers have genetically altered lignin, although, in a number of cases, these efforts have resulted in an undesirable yield penalty. Recent findings have shown that by knocking out two subunits (MED5A and MED5B) of the transcriptional regulatory complex Mediator, the stunted growth phenotype of mutants in p-coumaroyl shikimate 3'-hydroxylase, reduced epidermal fluorescence 8-1 (ref8-1), can be alleviated. Furthermore, these plants synthesize a lignin polymer almost entirely derived from p-coumaryl alcohol. Plants deficient in cinnamyl alcohol dehydrogenase (CAD) are notable in that they primarily incorporate coniferaldehyde and sinapaldehyde into their lignin. We tested the hypothesis that by stacking mutations in the genes encoding for the CAD paralogs C and D on an Arabidopsis (Arabidopsis thaliana) med5a/5b ref8-1 genetic background, the biosynthesis of p-coumaryl alcohol would be blocked, making p-coumaraldehyde available for polymerization into a novel kind of lignin. The med5a/5b ref8-1 cadc cadd plants are viable, but lignin analysis demonstrated that they continue to synthesize p-hydroxyphenyl lignin despite being mutated for the CADs typically considered to be required for monolignol biosynthesis. In addition, enzyme activity tests showed that even in the absence of CADC and CADD, there is high CAD activity in stems. We tested the potential involvement of other CADs in p-coumaraldehyde biosynthesis in the quintuple mutant by mutating them using the CRISPR/Cas9 system. Lignin analysis demonstrated that the resulting hextuple mutant plants continue to deposit p-coumaryl alcohol-derived lignin, demonstrating a route for the synthesis of p-hydroxyphenyl lignin in Arabidopsis independent of four CAD isoforms.
© American Society of Plant Biologists 2022. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures
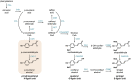


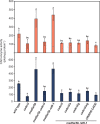
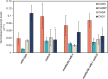

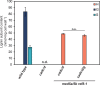

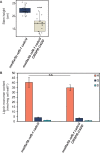
Comment in
-
Yet another twist in lignin biosynthesis: Is there a specific alcohol dehydrogenase for H-lignin production?Plant Physiol. 2022 Aug 1;189(4):1884-1886. doi: 10.1093/plphys/kiac249. Plant Physiol. 2022. PMID: 35639729 Free PMC article. No abstract available.
References
-
- Anderson NA, Tobimatsu Y, Ciesielski PN, Ximenes E, Ralph J, Donohoe BS, Ladisch M, Chapple C (2015) Manipulation of guaiacyl and syringyl monomer biosynthesis in an Arabidopsis cinnamyl alcohol dehydrogenase mutant results in atypical lignin biosynthesis and modified cell wall structure. Plant Cell 27: 2195–2209 - PMC - PubMed
-
- Bonawitz ND, Kim JI, Tobimatsu Y, Ciesielski PN, Anderson NA, Ximenes E, Maeda J, Ralph J, Donohoe BS, Ladisch M, et al. (2014) Disruption of Mediator rescues the stunted growth of a lignin-deficient Arabidopsis mutant. Nature 509: 376–380 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

