Upon heat stress processing of ribosomal RNA precursors into mature rRNAs is compromised after cleavage at primary P site in Arabidopsis thalian a
- PMID: 35522061
- PMCID: PMC9090299
- DOI: 10.1080/15476286.2022.2071517
Upon heat stress processing of ribosomal RNA precursors into mature rRNAs is compromised after cleavage at primary P site in Arabidopsis thalian a
Abstract
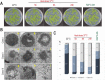
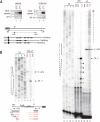
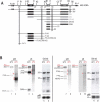
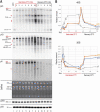
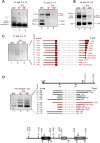
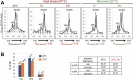
Transcription and processing of 45S rRNAs in the nucleolus are keystones of ribosome biogenesis. While these processes are severely impacted by stress conditions in multiple species, primarily upon heat exposure, we lack information about the molecular mechanisms allowing sessile organisms without a temperature-control system, like plants, to cope with such circumstances. We show that heat stress disturbs nucleolar structure, inhibits pre-rRNA processing and provokes imbalanced ribosome profiles in Arabidopsis thaliana plants. Notably, the accuracy of transcription initiation and cleavage at the primary P site in the 5'ETS (5' External Transcribed Spacer) are not affected but the levels of primary 45S and 35S transcripts are, respectively, increased and reduced. In contrast, precursors of 18S, 5.8S and 25S RNAs are rapidly undetectable upon heat stress. Remarkably, nucleolar structure, pre-rRNAs from major ITS1 processing pathway and ribosome profiles are restored after returning to optimal conditions, shedding light on the extreme plasticity of nucleolar functions in plant cells. Further genetic and molecular analysis to identify molecular clues implicated in these nucleolar responses indicate that cleavage rate at P site and nucleolin protein expression can act as a checkpoint control towards a productive pre-rRNA processing pathway.
Keywords: Arabidopsis; Nucleolus; heat stress; rRNA processing; ribosome.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures

References
-
- Sharma S, Lafontaine DL.. ‘view from a bridge’: a new perspective on eukaryotic rRNA base modification. Trends Biochem Sci. 2015;40(10):560–575. - PubMed
-
- Tomecki R, Sikorski PJ, Zakrzewska-Placzek M. Comparison of preribosomal RNA processing pathways in yeast, plant and human cells - focus on coordinated action of endo- and exoribonucleases. FEBS Lett. 2017;591(13):1801–1850. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
