A novel gene from the acidophilic bacterium Leptospirillum sp. CF-1 and its role in oxidative stress and chromate tolerance
- PMID: 35525996
- PMCID: PMC9080137
- DOI: 10.1186/s40659-022-00388-0
A novel gene from the acidophilic bacterium Leptospirillum sp. CF-1 and its role in oxidative stress and chromate tolerance
Abstract
Background: Acidophilic microorganisms like Leptospirillum sp. CF-1 thrive in environments with extremely low pH and high concentrations of dissolved heavy metals that can induce the generation of reactive oxygen species (ROS). Several hypothetical genes and proteins from Leptospirillum sp. CF-1 are known to be up-regulated under oxidative stress conditions.
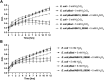
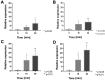
Results: In the present work, the function of hypothetical gene ABH19_09590 from Leptospirillum sp. CF-1 was studied. Heterologous expression of this gene in Escherichia coli led to an increase in the ability to grow under oxidant conditions with 5 mM K2CrO4 or 5 mM H2O2. Similarly, a significant reduction in ROS production in E. coli transformed with a plasmid carrying ABH19_09590 was observed after exposure to these oxidative stress elicitors for 30 min, compared to a strain complemented with the empty vector. A co-transcriptional study using RT-PCR showed that ABH19_09590 is contained in an operon, here named the "och" operon, that also contains ABH19_09585, ABH19_09595 and ABH19_09600 genes. The expression of the och operon was significantly up-regulated in Leptospirillum sp. CF-1 exposed to 5 mM K2CrO4 for 15 and 30 min. Genes of this operon potentially encode a NADH:ubiquinone oxidoreductase, a CXXC motif-containing protein likely involved in thiol/disulfide exchange, a hypothetical protein, and a di-hydroxy-acid dehydratase. A comparative genomic analysis revealed that the och operon is a characteristic genetic determinant of the Leptospirillum genus that is not present in other acidophiles.
Conclusions: Altogether, these results suggest that the och operon plays a protective role against chromate and hydrogen peroxide and is an important mechanism required to face polyextremophilic conditions in acid environments.
Keywords: Chromate; Hydrogen peroxide; Hypothetical proteins; Leptospirillum; Och operon; Oxidative stress.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Jones GC, Corin KC, Van Hille RP, Harrison STL. The generation of toxic reactive oxygen species (ROS) from mechanically activated sulphide concentrates and its effect on thermophilic bioleaching. Miner Eng. 2011;24(11):1198–1208. doi: 10.1016/j.mineng.2011.05.016. - DOI
-
- Christel S, Herold M, Bellenberg S, El Hajjami M, Buetti-Dinh A, Pivkin IV, Sand W, Wilmes P, Poetsch A, Dopson M. Multi-omics reveals the lifestyle of the acidophilic, mineral-oxidizing model species Leptospirillum ferriphilum. Appl Environ Microbiol. 2018;84:e02091–e2117. doi: 10.1128/AEM.02091-17. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

