Identifying Immunological and Clinical Predictors of COVID-19 Severity and Sequelae by Mathematical Modeling
- PMID: 35529862
- PMCID: PMC9067542
- DOI: 10.3389/fimmu.2022.865845
Identifying Immunological and Clinical Predictors of COVID-19 Severity and Sequelae by Mathematical Modeling
Abstract
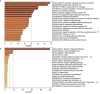
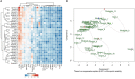
Since its emergence as a pandemic in March 2020, coronavirus disease (COVID-19) outcome has been explored via several predictive models, using specific clinical or biochemical parameters. In the current study, we developed an integrative non-linear predictive model of COVID-19 outcome, using clinical, biochemical, immunological, and radiological data of patients with different disease severities. Initially, the immunological signature of the disease was investigated through transcriptomics analysis of nasopharyngeal swab samples of patients with different COVID-19 severity versus control subjects (exploratory cohort, n=61), identifying significant differential expression of several cytokines. Accordingly, 24 cytokines were validated using a multiplex assay in the serum of COVID-19 patients and control subjects (validation cohort, n=77). Predictors of severity were Interleukin (IL)-10, Programmed Death-Ligand-1 (PDL-1), Tumor necrosis factors-α, absolute neutrophil count, C-reactive protein, lactate dehydrogenase, blood urea nitrogen, and ferritin; with high predictive efficacy (AUC=0.93 and 0.98 using ROC analysis of the predictive capacity of cytokines and biochemical markers, respectively). Increased IL-6 and granzyme B were found to predict liver injury in COVID-19 patients, whereas interferon-gamma (IFN-γ), IL-1 receptor-a (IL-1Ra) and PD-L1 were predictors of remarkable radiological findings. The model revealed consistent elevation of IL-15 and IL-10 in severe cases. Combining basic biochemical and radiological investigations with a limited number of curated cytokines will likely attain accurate predictive value in COVID-19. The model-derived cytokines highlight critical pathways in the pathophysiology of the COVID-19 with insight towards potential therapeutic targets. Our modeling methodology can be implemented using new datasets to identify key players and predict outcomes in new variants of COVID-19.
Keywords: Aritficial Intelligence; COVID-19; Machine Learning; RNA seq; ROC analysis; multiplex; transcriptomics.
Copyright © 2022 Elemam, Hammoudeh, Salameh, Mahboub, Alsafar, Talaat, Habib, Siddiqui, Hassan, Al-Assaf, Taneera, Sulaiman, Hamoudi, Maghazachi, Hamid and Saber-Ayad.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

