Identification of Bioactive Components of Stephania epigaea Lo and Their Potential Therapeutic Targets by UPLC-MS/MS and Network Pharmacology
- PMID: 35529936
- PMCID: PMC9068296
- DOI: 10.1155/2022/3641586
Identification of Bioactive Components of Stephania epigaea Lo and Their Potential Therapeutic Targets by UPLC-MS/MS and Network Pharmacology
Abstract
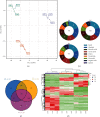
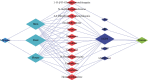
Stephania epigaea, an important traditional folk medicinal plant, elucidating its bioactive compound profiles and their molecular mechanisms of action on human health, would better understand its traditional therapies and guide their use in preclinical and clinical. This study aims to detect the critical therapeutic compounds, predict their targets, and explore potential therapeutic molecular mechanisms. This work first determined metabolites from roots, stems, and flowering twigs of S. epigaea by a widely targeted metabolomic analysis assay. Then, the drug likeness of the compounds and their pharmacokinetic profiles were screened by the ADMETlab server. The target proteins of active compounds were further analyzed by PPI combing with GO and KEGG cluster enrichment analysis. Finally, the interaction networks between essential compounds, targets, and disease-associated pathways were constructed, and the essential compounds binding to their possible target proteins were verified by molecular docking. Five key target proteins (EGFR, HSP90AA1, SRC, TNF, and CASP3) and twelve correlated metabolites, including aknadinine, cephakicine, homostephanoline, and N-methylliriodendronine associated with medical applications of S. epigaea, were identified, and the compounds and protein interactions were verified. The key active ingredients are mainly accumulated in the root, which indicates that the root is the main medicinal tissue. This study demonstrated that S. epigaea might exert the desired disease efficacy mainly through twelve components interacting via five essential target proteins. EGFR is the most critical one, which deserves further verification by biological studies.
Copyright © 2022 Xingyu Li et al.
Conflict of interest statement
The authors declare that there are no conflicts of interest regarding the publication of this paper.
Figures




References
-
- Wu Z., Raven P. H., Hong D., Missouri Botanical Garden . Flora of China . Vol. 1. Beijing, China: Science Press; 1996.
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

