Investigation of the effects of overexpression of jumping translocation breakpoint (JTB) protein in MCF7 cells for potential use as a biomarker in breast cancer
- PMID: 35530281
- PMCID: PMC9077082
Investigation of the effects of overexpression of jumping translocation breakpoint (JTB) protein in MCF7 cells for potential use as a biomarker in breast cancer
Abstract
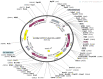
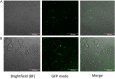
Jumping translocation breakpoint (JTB) gene acts as a tumor suppressor or an oncogene in different malignancies, including breast cancer (BC), where it was reported as overexpressed. However, the molecular functions, biological processes and underlying mechanisms through which JTB protein causes increased cell growth, proliferation and invasion is still not fully deciphered. Our goal is to identify the functions of JTB protein by cellular proteomics approaches. MCF7 breast cancer cells were transfected with sense orientation of hJTB cDNA in HA, His and FLAG tagged CMV expression vector to overexpress hJTB and the expression levels were confirmed by Western blotting (WB). Proteins extracted from transfected cells were separated by SDS-PAGE and the in-gel digested peptides were analyzed by nano-liquid chromatography tandem mass spectrometry (nanoLC-MS/MS). By comparing the proteome of cells with upregulated conditions of JTB vs control and identifying the protein dysregulation patterns, we aim to understand the function of this protein and its contribution to tumorigenesis. Gene Set Enrichment Analysis (GSEA) algorithm was performed to investigate the biological processes and pathways that are associated with the JTB protein upregulation. The results demonstrated four significantly enriched gene sets from the following significantly upregulated pathways: mitotic spindle assembly, estrogen response late, epithelial-to-mesenchymal transition (EMT) and estrogen response early. JTB protein itself is involved in mitotic spindle pathway by its role in cell division/cytokinesis, and within estrogen response early and late pathways, contributing to discrimination between luminal and mesenchymal breast cancer. Thus, the overexpressed JTB condition was significantly associated with an increased expression of ACTNs, FLNA, FLNB, EZR, MYOF, COL3A1, COL11A1, HSPA1A, HSP90A, WDR, EPPK1, FASN and FOXA1 proteins related to deregulation of cytoskeletal organization and biogenesis, mitotic spindle organization, ECM remodeling, cellular response to estrogen, proliferation, migration, metastasis, increased lipid biogenesis, endocrine therapy resistance, antiapoptosis and discrimination between different breast cancer subtypes. Other upregulated proteins for overexpressed JTB condition are involved in multiple cellular functions and pathways that become dysregulated, such as tumor microenvironment (TME) acidification, the transmembrane transport pathways, glycolytic flux, iron metabolism and oxidative stress, metabolic reprogramming, nucleocytosolic mRNA transport, transcriptional activation, chromatin remodeling, modulation of cell death pathways, stress responsive pathways, and cancer drug resistance. The downregulated proteins for overexpressed JTB condition are involved in adaptive communication between external and internal environment of cells and maintenance between pro-apoptotic and anti-apoptotic signaling pathways, vesicle trafficking and secretion, DNA lesions repair and suppression of genes involved in tumor progression, proteostasis, redox state regulation, biosynthesis of macromolecules, lipolytic pathway, carbohydrate metabolism, dysregulation of ubiquitin-mediated degradation system, cancer cell immune escape, cell-to-cell and cell-to-ECM interactions, and cytoskeletal behaviour. There were no significantly enriched downregulated pathways.
Keywords: Breast cancer; JTB overexpressed condition; jumping translocation breakpoint (JTB) protein; proteomics.
AJCR Copyright © 2022.
Conflict of interest statement
None.
Figures



References
-
- Ghoncheh M, Pournamdar Z, Salehiniya H. Incidence and mortality and epidemiology of breast cancer in the world. Asian Pac J Cancer Prev. 2016;17:43–46. - PubMed
-
- DeSantis CE, Ma J, Gaudet MM, Newman LA, Miller KD, Goding Sauer A, Jemal A, Siegel RL. Breast cancer statistics, 2019. CA Cancer J Clin. 2019;69:438–451. - PubMed
-
- Hatakeyama S, Osawa M, Omine M, Ishikawa F. JTB: a novel membrane protein gene at 1q21 rearranged in a jumping translocation. Oncogene. 1999;18:2085–2090. - PubMed
-
- Kanome T, Itoh N, Ishikawa F, Mori K, Kim-Kaneyama JR, Nose K, Shibanuma M. Characterization of jumping translocation breakpoint (JTB) gene product isolated as a TGF-β1-inducible clone involved in regulation of mitochondrial function, cell growth and cell death. Oncogene. 2007;26:5991–6001. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
