Dynamic proteomic change of tumor and immune organs in an immune-competent hepatocellular carcinoma mouse model
- PMID: 35530287
- PMCID: PMC9077077
Dynamic proteomic change of tumor and immune organs in an immune-competent hepatocellular carcinoma mouse model
Abstract
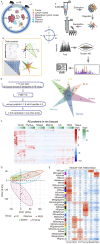
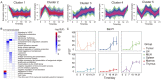
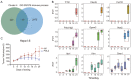
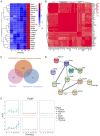
Subcutaneous implantation of a human cancer cell line in immune-deficient mice (CDX) is a commonly used tool in preclinical studies for the assessment of potential anti-cancer drugs. As immunotherapy is transforming cancer treatment, tumor models in immunocompetent mice are necessary for us to understand the immune aspects of tumor biology. However, the systemic immune response to the implantation of cancer cells at proteome level is unclear. In this study, we characterized the dynamic proteomic changes of subcutaneous tumors and 5 immune organs (draining lymph node, mesenteric lymph node, spleen, thymus and marrow) at six time points after implantation using a Hepa1-6 derived allograft mouse model. Our data suggest that interaction of the implanted tumor cells with mouse immune system followed the trajectory of "tumor rejection" to "immune evasion" in that the tumor gained the ability to evade the immune system for growth. Furthermore, anti-PDL2 antibody was validated here as an optional immunotherapy strategy to inhibit the growth of Hepa1-6 subcutaneous tumors. These findings from our study provided valuable information for the understanding of tumor and immune interaction and shed light on the rational design for clinical cancer treatment and other preclinical experiments.
Keywords: Hepa1-6; PDL2; allograft mouse model; proteomics.
AJCR Copyright © 2022.
Conflict of interest statement
None.
Figures
References
-
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674. - PubMed
-
- Galon J, Bruni D. Tumor immunology and tumor evolution: intertwined histories. Immunity. 2020;52:55–81. - PubMed
-
- Whiteside TL, Mandapathil M, Szczepanski M, Szajnik M. Mechanisms of tumor escape from the immune system: adenosine-producing Treg, exosomes and tumor-associated TLRs. Bull Cancer. 2011;98:E25–31. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
