Profiling the Human Phosphoproteome to Estimate the True Extent of Protein Phosphorylation
- PMID: 35532924
- PMCID: PMC9171898
- DOI: 10.1021/acs.jproteome.2c00131
Profiling the Human Phosphoproteome to Estimate the True Extent of Protein Phosphorylation
Abstract
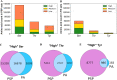
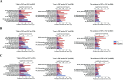
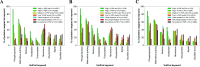
Public phosphorylation databases such as PhosphoSitePlus (PSP) and PeptideAtlas (PA) compile results from published papers or openly available mass spectrometry (MS) data. However, there is no database-level control for false discovery of sites, likely leading to the overestimation of true phosphosites. By profiling the human phosphoproteome, we estimate the false discovery rate (FDR) of phosphosites and predict a more realistic count of true identifications. We rank sites into phosphorylation likelihood sets and analyze them in terms of conservation across 100 species, sequence properties, and functional annotations. We demonstrate significant differences between the sets and develop a method for independent phosphosite FDR estimation. Remarkably, we report estimated FDRs of 84, 98, and 82% within sets of phosphoserine (pSer), phosphothreonine (pThr), and phosphotyrosine (pTyr) sites, respectively, that are supported by only a single piece of identification evidence─the majority of sites in PSP. We estimate that around 62 000 Ser, 8000 Thr, and 12 000 Tyr phosphosites in the human proteome are likely to be true, which is lower than most published estimates. Furthermore, our analysis estimates that 86 000 Ser, 50 000 Thr, and 26 000 Tyr phosphosites are likely false-positive identifications, highlighting the significant potential of false-positive data to be present in phosphorylation databases.
Keywords: PeptideAtlas; PhosphoSitePlus; UniProt; database; evolutionary conservation; false discovery rate; mass spectrometry; phosphopeptides; phosphoproteomics; phosphorylation; phosphosites; proteome; proteomics.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
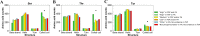
Similar articles
-
Phosphoproteome analysis of E. coli reveals evolutionary conservation of bacterial Ser/Thr/Tyr phosphorylation.Mol Cell Proteomics. 2008 Feb;7(2):299-307. doi: 10.1074/mcp.M700311-MCP200. Epub 2007 Oct 15. Mol Cell Proteomics. 2008. PMID: 17938405
-
Phosphorylation in the Plasmodium falciparum Proteome: A Meta-Analysis of Publicly Available Data Sets.J Proteome Res. 2024 Dec 6;23(12):5326-5341. doi: 10.1021/acs.jproteome.4c00418. Epub 2024 Oct 30. J Proteome Res. 2024. PMID: 39475123 Free PMC article.
-
High-Throughput Characterization of Histidine Phosphorylation Sites Using UPAX and Tandem Mass Spectrometry.Methods Mol Biol. 2020;2077:225-235. doi: 10.1007/978-1-4939-9884-5_15. Methods Mol Biol. 2020. PMID: 31707662
-
Improving Phosphoproteomics Profiling Using Data-Independent Mass Spectrometry.J Proteome Res. 2022 Aug 5;21(8):1789-1799. doi: 10.1021/acs.jproteome.2c00172. Epub 2022 Jul 25. J Proteome Res. 2022. PMID: 35877786 Review.
-
Targeted mass spectrometry: An emerging powerful approach to unblock the bottleneck in phosphoproteomics.J Chromatogr B Analyt Technol Biomed Life Sci. 2017 Jun 15;1055-1056:29-38. doi: 10.1016/j.jchromb.2017.04.026. Epub 2017 Apr 17. J Chromatogr B Analyt Technol Biomed Life Sci. 2017. PMID: 28441545 Review.
Cited by
-
Identifying and evaluating understudied protein kinases using biological and chemical criteria.RSC Med Chem. 2025 Jun 5;16(8):3386-92. doi: 10.1039/d5md00306g. Online ahead of print. RSC Med Chem. 2025. PMID: 40510904 Review.
-
The fitness cost of spurious phosphorylation.EMBO J. 2024 Oct;43(20):4720-4751. doi: 10.1038/s44318-024-00200-7. Epub 2024 Sep 10. EMBO J. 2024. PMID: 39256561 Free PMC article.
-
High-throughput approaches for the identification of ribosome heterogeneity.Philos Trans R Soc Lond B Biol Sci. 2025 Mar 6;380(1921):20230381. doi: 10.1098/rstb.2023.0381. Epub 2025 Mar 6. Philos Trans R Soc Lond B Biol Sci. 2025. PMID: 40045778 Free PMC article. Review.
-
The fitness cost of spurious phosphorylation.bioRxiv [Preprint]. 2023 Oct 10:2023.10.08.561337. doi: 10.1101/2023.10.08.561337. bioRxiv. 2023. Update in: EMBO J. 2024 Oct;43(20):4720-4751. doi: 10.1038/s44318-024-00200-7. PMID: 37873463 Free PMC article. Updated. Preprint.
-
Borrelia PeptideAtlas: A proteome resource of common Borrelia burgdorferi isolates for Lyme research.Sci Data. 2024 Dec 2;11(1):1313. doi: 10.1038/s41597-024-04047-9. Sci Data. 2024. PMID: 39622905 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- C1443/A22095/CRUK_/Cancer Research UK/United Kingdom
- R24 GM127667/GM/NIGMS NIH HHS/United States
- BB/M012557/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/S018514/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/R000182/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

