The developmental stage of the medulloblastoma cell-of-origin restricts Sonic hedgehog pathway usage and drug sensitivity
- PMID: 35535520
- PMCID: PMC9234672
- DOI: 10.1242/jcs.258608
The developmental stage of the medulloblastoma cell-of-origin restricts Sonic hedgehog pathway usage and drug sensitivity
Abstract
Sonic hedgehog (SHH) medulloblastoma originates from the cerebellar granule neuron progenitor (CGNP) lineage, which depends on Hedgehog signaling for its perinatal expansion. Whereas SHH tumors exhibit overall deregulation of this pathway, they also show patient age-specific aberrations. To investigate whether the developmental stage of the CGNP can account for these age-specific lesions, we analyzed developing murine CGNP transcriptomes and observed highly dynamic gene expression as a function of age. Cross-species comparison with human SHH medulloblastoma showed partial maintenance of these expression patterns, and highlighted low primary cilium expression as hallmark of infant medulloblastoma and early embryonic CGNPs. This coincided with reduced responsiveness to upstream SHH pathway component Smoothened, whereas sensitivity to downstream components SUFU and GLI family proteins was retained. Together, these findings can explain the preference for SUFU mutations in infant medulloblastoma and suggest that drugs targeting the downstream SHH pathway will be most appropriate for infant patients.
Keywords: Cerebellar development; Cerebellar granule neuron progenitors; Medulloblastoma; Primary cilia; Sonic hedgehog signaling; Tumor cell-of-origin.
© 2022. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


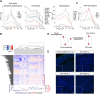
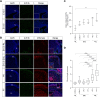
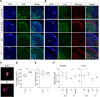

References
-
- Anne, S. L., Govek, E.-E., Ayrault, O., Kim, J. H., Zhu, X., Murphy, D. A., Van Aelst, L., Roussel, M. F. and Hatten, M. E. (2013). WNT3 inhibits cerebellar granule neuron progenitor proliferation and medulloblastoma formation via MAPK activation. PLoS One 8, e81769. 10.1371/journal.pone.0081769 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- UMCG Kanker Researchfonds (KRF)
- Stichting De Cock-Hadders
- 20151022024475/Indonesian Endowment Fund for Education (LPDP)
- Coordenacao de Aperfeicoamento de Pessoal de Nivel Superior
- University of Groningen and European Union
- Netherlands Organisation for Scientific Research (NWO)
- Stichting vrienden Beatrix Kinderziekenshuis
- RUG 2014-6903/KWF Kankerbestrijding
- UMCG Kanker Researchfonds
- Coordenação de Aperfeiçoamento de Pessoal de Nível Superior
- University of Groningen
- European Union
- Netherlands Organisation for Scientific Research
- Stichting Vrienden Beatrix Kinderziekenhuis
- Universitair Medisch Centrum Groningen
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

