Krüppel-like Factor 4 Supports the Expansion of Leukemia Stem Cells in MLL-AF9-driven Acute Myeloid Leukemia
- PMID: 35535819
- PMCID: PMC9406610
- DOI: 10.1093/stmcls/sxac033
Krüppel-like Factor 4 Supports the Expansion of Leukemia Stem Cells in MLL-AF9-driven Acute Myeloid Leukemia
Abstract
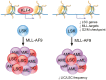
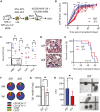
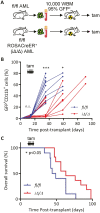
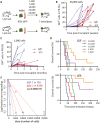
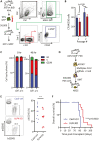
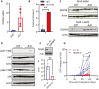
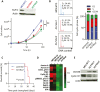
Acute myeloid leukemia (AML) is an aggressive malignancy of the bone marrow with 5-year overall survival of less than 10% in patients over the age of 65. Limited progress has been made in the patient outcome because of the inability to selectively eradicate the leukemic stem cells (LSC) driving the refractory and relapsed disease. Herein, we investigated the role of the reprogramming factor KLF4 in AML because of its critical role in the self-renewal and stemness of embryonic and cancer stem cells. Using a conditional Cre-lox Klf4 deletion system and the MLL-AF9 retroviral mouse model, we demonstrated that loss-of-KLF4 does not significantly affect the induction of leukemia but markedly decreased the frequency of LSCs evaluated in limiting-dose transplantation studies. Loss of KLF4 in leukemic granulocyte-macrophage progenitors (L-GMP), a population enriched for AML LSCs, showed lessened clonogenicity and percentage in the G2/M phase of the cell cycle. RNAseq analysis of purified L-GMPs revealed decreased expression of stemness genes and MLL-target genes and upregulation of the RNA sensing helicase DDX58. However, silencing of DDX58 in KLF4 knockout leukemia indicated that DDX58 is not mediating this phenotype. CRISPR/Cas9 deletion of KLF4 in MOLM13 cell line and AML patient-derived xenograft cells showed impaired expansion in vitro and in vivo associated with a defective G2/M checkpoint. Collectively, our data suggest a mechanism in which KLF4 promotes leukemia progression by establishing a gene expression profile in AML LSCs supporting cell division and stemness.
Keywords: KLF4; MLL-AF9; leukemic stem cells; transcription factor.
© The Author(s) 2022. Published by Oxford University Press. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures

Similar articles
-
Bone-Marrow-Targeted Nanocomposite Abrogates C-Myb-Survivin Cross Talk in MLL-AF9-Rearranged Acute Myeloid Leukemia in In Vitro and In Vivo Patient-Derived Xenograft Models.ACS Appl Mater Interfaces. 2025 Jan 8;17(1):711-724. doi: 10.1021/acsami.4c18737. Epub 2024 Dec 22. ACS Appl Mater Interfaces. 2025. PMID: 39711012
-
SUV39H1 regulates the progression of MLL-AF9-induced acute myeloid leukemia.Oncogene. 2020 Dec;39(50):7239-7252. doi: 10.1038/s41388-020-01495-6. Epub 2020 Oct 9. Oncogene. 2020. PMID: 33037410 Free PMC article.
-
Atg5-dependent autophagy contributes to the development of acute myeloid leukemia in an MLL-AF9-driven mouse model.Cell Death Dis. 2016 Sep 8;7(9):e2361. doi: 10.1038/cddis.2016.264. Cell Death Dis. 2016. PMID: 27607576 Free PMC article.
-
Exploring the contribution of Zfp521/ZNF521 on primary hematopoietic stem/progenitor cells and leukemia progression.Cell Tissue Res. 2024 Dec;398(3):161-173. doi: 10.1007/s00441-024-03926-2. Epub 2024 Oct 22. Cell Tissue Res. 2024. PMID: 39436449 Free PMC article. Review.
-
Learning from mouse models of MLL fusion gene-driven acute leukemia.Biochim Biophys Acta Gene Regul Mech. 2020 Aug;1863(8):194550. doi: 10.1016/j.bbagrm.2020.194550. Epub 2020 Apr 19. Biochim Biophys Acta Gene Regul Mech. 2020. PMID: 32320749 Review.
Cited by
-
The reprogramming factor KLF4 in normal and malignant blood cells.Front Immunol. 2025 Jun 16;16:1584181. doi: 10.3389/fimmu.2025.1584181. eCollection 2025. Front Immunol. 2025. PMID: 40589762 Free PMC article. Review.
-
KLF4-mediated upregulation of the NKG2D ligand MICA in acute myeloid leukemia: a novel therapeutic target identified by enChIP.Cell Commun Signal. 2023 May 4;21(1):94. doi: 10.1186/s12964-023-01118-z. Cell Commun Signal. 2023. PMID: 37143070 Free PMC article.
-
SWI/SNF Blockade Disrupts PU.1-Directed Enhancer Programs in Normal Hematopoietic Cells and Acute Myeloid Leukemia.Cancer Res. 2023 Apr 4;83(7):983-996. doi: 10.1158/0008-5472.CAN-22-2129. Cancer Res. 2023. PMID: 36662812 Free PMC article.
-
Role of the bone marrow vascular niche in chemotherapy for MLL-AF9-induced acute myeloid leukemia.Blood Sci. 2023 Apr 19;5(2):92-100. doi: 10.1097/BS9.0000000000000158. eCollection 2023 Apr. Blood Sci. 2023. PMID: 37228781 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

