Genome-wide identification and expression characterization of the DoG gene family of moso bamboo (Phyllostachys edulis)
- PMID: 35538420
- PMCID: PMC9092881
- DOI: 10.1186/s12864-022-08551-3
Genome-wide identification and expression characterization of the DoG gene family of moso bamboo (Phyllostachys edulis)
Erratum in
-
Correction: Genome-wide identification and expression characterization of the DoG gene family of moso bamboo (Phyllostachys edulis).BMC Genomics. 2023 Mar 17;24(1):128. doi: 10.1186/s12864-023-09130-w. BMC Genomics. 2023. PMID: 36932336 Free PMC article. No abstract available.
Abstract
Background: The DoG (Delay of Germination1) family plays a key regulatory role in seed dormancy and germination. However, to date, there is no complete genomic overview of the DoG gene family of any economically valuable crop, including moso bamboo (Phyllostachys edulis), and no studies have been conducted to characterize its expression profile. To identify the DoG gene members of moso bamboo (PeDoG) and to investigate their family structural features and tissue expression profile characteristics, a study was conducted. Based on the whole genome and differential transcriptome data, in this investigation, we have scrutinized the physicochemical properties, gene structure, cis-acting elements, phylogenetic relationships, conserved structural (CS) domains, CS motifs and expression patterns of the PeDoG1 family of moso bamboo.
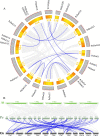
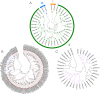
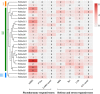
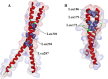
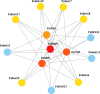
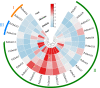
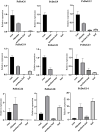
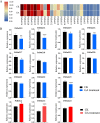
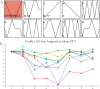
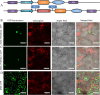
Results: The DoG family genes of moso bamboo were found distributed across 16 chromosomal scaffolds with 24 members. All members were found to carry DoG1 structural domains, while 23 members additionally possessed basic leucine zipper (bZIP) structural domains. We could divide the PeDoG genes into three subfamilies based on phylogenetic relationships. Covariance analysis revealed that tandem duplication was the main driver of amplification of the PeDoG genes. The upstream promoter of these genes containing several cis-acting elements indicates a plausible role in abiotic stress and hormone induction. Gene expression pattern according to transcriptome data revealed participation of the PeDoG genes in tissue and organ development. Analysis using Short Time-series Expression Miner (STEM) tool revealed that the PeDoG gene family is also associated with rapid early shoot growth. Gene ontology (GO) and KEGG analyses showed a dual role of the PeDoG genes. We found that PeDoGs has a possible role as bZIP transcription factors by regulating Polar like1 (PL1) gene expression, and thereby playing a disease response role in moso bamboo. Quantitative gene expression of the PeDoG genes revealed that they were abundantly expressed in roots and leaves, and could be induced in response to gibberellin (GA).
Conclusion: In this study, we found that the PeDoG genes are involved in a wide range of activities such as growth and development, stress response and transcription. This forms the first report of PeDoG genes and their potential roles in moso bamboo.
Keywords: BZIP transcription factor; Bioinformatics; DoG gene family; Gene expression; Phyllostachys edulis.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Genome-wide analysis of the KNOX gene family in Moso bamboo: insights into their role in promoting the rapid shoot growth.BMC Plant Biol. 2024 Mar 25;24(1):213. doi: 10.1186/s12870-024-04883-2. BMC Plant Biol. 2024. PMID: 38528453 Free PMC article.
-
Genome-wide identification and expression analysis of LBD transcription factor genes in Moso bamboo (Phyllostachys edulis).BMC Plant Biol. 2021 Jun 28;21(1):296. doi: 10.1186/s12870-021-03078-3. BMC Plant Biol. 2021. PMID: 34182934 Free PMC article.
-
Genome-Wide Identification and Expression Analyses of the bZIP Transcription Factor Genes in moso bamboo (Phyllostachys edulis).Int J Mol Sci. 2019 May 5;20(9):2203. doi: 10.3390/ijms20092203. Int J Mol Sci. 2019. PMID: 31060272 Free PMC article.
-
Transcriptomic Complexity of Culm Growth and Development in Different Types of Moso Bamboo.Int J Mol Sci. 2023 Apr 18;24(8):7425. doi: 10.3390/ijms24087425. Int J Mol Sci. 2023. PMID: 37108588 Free PMC article. Review.
-
Unraveling the intricate tapestry of bamboo transcription factors in abiotic stress signaling and resilience with special reference to moso bamboo family.Biochim Biophys Acta Gen Subj. 2025 Feb;1869(2):130755. doi: 10.1016/j.bbagen.2024.130755. Epub 2024 Dec 29. Biochim Biophys Acta Gen Subj. 2025. PMID: 39740732 Review.
Cited by
-
Unraveling the Guardians of Growth: A Comprehensive Analysis of the Aux/IAA and ARF Gene Families in Populus simonii.Plants (Basel). 2023 Oct 13;12(20):3566. doi: 10.3390/plants12203566. Plants (Basel). 2023. PMID: 37896029 Free PMC article.
-
Genome-wide identification and expression characterization of the GH3 gene family of tea plant (Camellia sinensis).BMC Genomics. 2024 Jan 27;25(1):120. doi: 10.1186/s12864-024-10004-y. BMC Genomics. 2024. PMID: 38280985 Free PMC article.
-
Evolutionary and functional analysis of the DIR gene family in Moso bamboo: insights into rapid shoot growth and stress responses.Front Plant Sci. 2025 Feb 25;16:1535733. doi: 10.3389/fpls.2025.1535733. eCollection 2025. Front Plant Sci. 2025. PMID: 40070714 Free PMC article.
-
Genome-wide identification and expression analysis of the WRKY transcription factors related to sesquiterpenes biosynthesis in Atractylodes lancea.Front Genet. 2025 May 15;16:1551991. doi: 10.3389/fgene.2025.1551991. eCollection 2025. Front Genet. 2025. PMID: 40444277 Free PMC article.
-
Correction: Genome-wide identification and expression characterization of the DoG gene family of moso bamboo (Phyllostachys edulis).BMC Genomics. 2023 Mar 17;24(1):128. doi: 10.1186/s12864-023-09130-w. BMC Genomics. 2023. PMID: 36932336 Free PMC article. No abstract available.
References
-
- Ramakrishnan M, Yrjälä K, Vinod KK, Sharma A, Cho J, Satheesh V, Zhou M. Genetics and genomics of moso bamboo (Phyllostachys edulis): Current status, future challenges, and biotechnological opportunities toward a sustainable bamboo industry. Food and Energy Security. 2020;9(4):299. doi: 10.1002/fes3.229. - DOI
-
- Qiao G, Li H, Liu M, Jiang J, Yin Y, Zhang L, Zhuo R. Callus induction and plant regeneration from anthers of Dendrocalamus latiflorus Munro. In Vitro Cellular & Developmental Biology - Plant. 2013;49(4):375–382. doi: 10.1007/s11627-013-9498-8. - DOI
-
- Zhu Zh, Wei J. Sustainable Bamboo Development. Oxfordshire, UK: CABI; 2018. p. 103.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

