Occurrence and Nature of Off-Target Modifications by CRISPR-Cas Genome Editing in Plants
- PMID: 35548699
- PMCID: PMC9075866
- DOI: 10.1021/acsagscitech.1c00270
Occurrence and Nature of Off-Target Modifications by CRISPR-Cas Genome Editing in Plants
Abstract
CRISPR-Cas-based genome editing allows for precise and targeted genetic modification of plants. Nevertheless, unintended off-target edits can arise that might confer risks when present in gene-edited food crops. Through an extensive literature review we gathered information on CRISPR-Cas off-target edits in plants. Most observed off-target changes were small insertions or deletions (1-22 bp) or nucleotide substitutions, and large deletions (>100 bp) were rare. One study detected the insertion of vector-derived DNA sequences, which is important considering the risk assessment of gene-edited plants. Off-target sites had few mismatches (1-3 nt) with the target sequence and were mainly located in protein-coding regions, often in target gene homologues. Off-targets edits were predominantly detected via biased analysis of predicted off-target sites instead of unbiased genome-wide analysis. CRISPR-Cas-edited plants showed lower off-target mutation frequencies than conventionally bred plants. This Review can aid discussions on the relevance of evaluating off-target modifications for risk assessment of CRISPR-Cas-edited plants.
© 2022 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
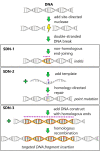
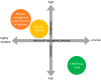
References
Publication types
LinkOut - more resources
Full Text Sources
Research Materials
