GABPA-activated TGFBR2 transcription inhibits aggressiveness but is epigenetically erased by oncometabolites in renal cell carcinoma
- PMID: 35549739
- PMCID: PMC9097325
- DOI: 10.1186/s13046-022-02382-6
GABPA-activated TGFBR2 transcription inhibits aggressiveness but is epigenetically erased by oncometabolites in renal cell carcinoma
Abstract
Background: The ETS transcription factor GABPA has long been thought of as an oncogenic factor and recently suggested as a target for cancer therapy due to its critical effect on telomerase activation, but the role of GABPA in clear cell renal cell carcinoma (ccRCC) is unclear. In addition, ccRCC is characterized by metabolic reprograming with aberrant accumulation of L-2-hydroxyglurate (L-2HG), an oncometabolite that has been shown to promote ccRCC development and progression by inducing DNA methylation, however, its downstream effectors remain poorly defined.
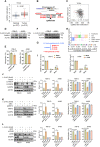
Methods: siRNAs and expression vectors were used to manipulate the expression of GABPA and other factors and to determine cellular/molecular and phenotypic alterations. RNA sequencing and ChIP assays were performed to identify GABPA target genes. A human ccRCC xenograft model in mice was used to evaluate the effect of GABPA overexpression on in vivo tumorigenesis and metastasis. ccRCC cells were incubated with L-2-HG to analyze GABPA expression and methylation. We carried out immunohistochemistry on patient specimens and TCGA dataset analyses to assess the effect of GABPA on ccRCC survival.
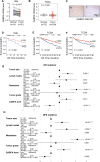
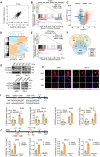
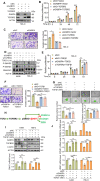
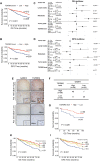
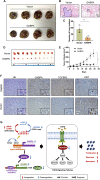
Results: GABPA depletion, although inhibiting telomerase expression, robustly enhanced proliferation, invasion and stemness of ccRCC cells, whereas GABPA overexpression exhibited opposite effects, strongly inhibiting in vivo metastasis and carcinogenesis. TGFBR2 was identified as the GABPA target gene through which GABPA governed the TGFβ signaling to dictate ccRCC phenotypes. GABPA and TGFBR2 phenocopies each other in ccRCC cells. Higher GABPA or TGFBR2 expression predicted longer survival in patients with ccRCC. Incubation of ccRCC cells with L-2-HG mimics GABPA-knockdown-mediated phenotypic alterations. L-2-HG silenced the expression of GABPA in ccRCC cells by increasing its methylation.
Conclusions: GABPA acts as a tumor suppressor by stimulating TGFBR2 expression and TGFβ signaling, while L-2-HG epigenetically inhibits GABPA expression, disrupting the GABPA-TGFβ loop to drive ccRCC aggressiveness. These results exemplify how oncometabolites erase tumor suppressive function for cancer development/progression. Restoring GABPA expression using DNA methylation inhibitors or other approaches, rather than targeting it, may be a novel strategy for ccRCC therapy.
Keywords: GABPA; L-2-HG; Oncometabolite; TGFBR2; ccRCC.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
GABPA inhibits tumorigenesis in clear cell renal cell carcinoma by regulating ferroptosis through ACSL4.Sci Rep. 2024 Nov 3;14(1):26521. doi: 10.1038/s41598-024-78441-z. Sci Rep. 2024. PMID: 39489850 Free PMC article.
-
GABPA is a master regulator of luminal identity and restrains aggressive diseases in bladder cancer.Cell Death Differ. 2020 Jun;27(6):1862-1877. doi: 10.1038/s41418-019-0466-7. Epub 2019 Dec 4. Cell Death Differ. 2020. PMID: 31802036 Free PMC article.
-
GABPA inhibits invasion/metastasis in papillary thyroid carcinoma by regulating DICER1 expression.Oncogene. 2019 Feb;38(7):965-979. doi: 10.1038/s41388-018-0483-x. Epub 2018 Sep 4. Oncogene. 2019. PMID: 30181547
-
TERT promoter mutations and GABP transcription factors in carcinogenesis: More foes than friends.Cancer Lett. 2020 Nov 28;493:1-9. doi: 10.1016/j.canlet.2020.07.003. Epub 2020 Aug 6. Cancer Lett. 2020. PMID: 32768523 Review.
-
Oncometabolites in renal cancer.Nat Rev Nephrol. 2020 Mar;16(3):156-172. doi: 10.1038/s41581-019-0210-z. Epub 2019 Oct 21. Nat Rev Nephrol. 2020. PMID: 31636445 Free PMC article. Review.
Cited by
-
The cuproptosis-associated 13 gene signature as a robust predictor for outcome and response to immune- and targeted-therapies in clear cell renal cell carcinoma.Front Immunol. 2022 Sep 5;13:971142. doi: 10.3389/fimmu.2022.971142. eCollection 2022. Front Immunol. 2022. PMID: 36131921 Free PMC article.
-
A regulatory network controlling ovarian granulosa cell death.Cell Death Discov. 2023 Feb 20;9(1):70. doi: 10.1038/s41420-023-01346-9. Cell Death Discov. 2023. PMID: 36806197 Free PMC article.
-
Thyroid carcinoma-featured telomerase activation and telomere maintenance: Biology and translational/clinical significance.Clin Transl Med. 2022 Nov;12(11):e1111. doi: 10.1002/ctm2.1111. Clin Transl Med. 2022. PMID: 36394204 Free PMC article. Review.
-
Abnormal changes in metabolites caused by m6A methylation modification: The leading factors that induce the formation of immunosuppressive tumor microenvironment and their promising potential for clinical application.J Adv Res. 2025 Apr;70:159-186. doi: 10.1016/j.jare.2024.04.016. Epub 2024 Apr 25. J Adv Res. 2025. PMID: 38677545 Free PMC article. Review.
-
Cuproptosis and its application in different cancers: an overview.Mol Cell Biochem. 2023 Dec;478(12):2683-2693. doi: 10.1007/s11010-023-04693-4. Epub 2023 Mar 13. Mol Cell Biochem. 2023. PMID: 36914880
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

