Towards Large-Scale Integrative Taxonomy (LIT): Resolving the Data Conundrum for Dark Taxa
- PMID: 35556139
- PMCID: PMC9558837
- DOI: 10.1093/sysbio/syac033
Towards Large-Scale Integrative Taxonomy (LIT): Resolving the Data Conundrum for Dark Taxa
Abstract
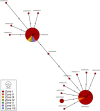
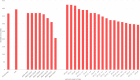
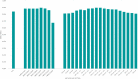
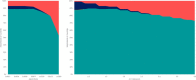
New, rapid, accurate, scalable, and cost-effective species discovery and delimitation methods are needed for tackling "dark taxa," here defined as groups for which $<$10$\%$ of all species are described and the estimated diversity exceeds 1,000 species. Species delimitation for these taxa should be based on multiple data sources ("integrative taxonomy") but collecting multiple types of data risks impeding a discovery process that is already too slow. We here develop large-scale integrative taxonomy (LIT), an explicit method where preliminary species hypotheses are generated based on inexpensive data that can be obtained quickly and cost-effectively. These hypotheses are then evaluated based on a more expensive type of "validation data" that is only obtained for specimens selected based on objective criteria applied to the preliminary species hypotheses. We here use this approach to sort 18,000 scuttle flies (Diptera: Phoridae) into 315 preliminary species hypotheses based on next-generation sequencing barcode (313 bp) clusters (using objective clustering [OC] with a 3$\%$ threshold). These clusters are then evaluated with morphology as the validation data. We develop quantitative indicators for predicting which barcode clusters are likely to be incongruent with morphospecies by randomly selecting 100 clusters for in-depth validation with morphology. A linear model demonstrates that the best predictors for incongruence between barcode clusters and morphology are maximum p-distance within the cluster and a newly proposed index that measures cluster stability across different clustering thresholds. A test of these indicators using the 215 remaining clusters reveals that these predictors correctly identify all clusters that are incongruent with morphology. In our study, all morphospecies are true or disjoint subsets of the initial barcode clusters so that all incongruence can be eliminated by varying clustering thresholds. This leads to a discussion of when a third data source is needed to resolve incongruent grouping statements. The morphological validation step in our study involved 1,039 specimens (5.8$\%$ of the total). The formal LIT protocol we propose would only have required the study of 915 (5.1$\%$: 2.5 specimens per species), as we show that clusters without signatures of incongruence can be validated by only studying two specimens representing the most divergent haplotypes. To test the generality of our results across different barcode clustering techniques, we establish that the levels of incongruence are similar across OC, Automatic Barcode Gap Discovery (ABGD), Poisson Tree Processes (PTP), and Refined Single Linkage (RESL) (used by Barcode of Life Data System to assign Barcode Index Numbers [BINs]). OC and ABGD achieved a maximum congruence score with the morphology of 89$\%$ while PTP was slightly less effective (84$\%$). RESL could only be tested for a subset of the specimens because the algorithm is not public. BINs based on 277 of the original 1,714 haplotypes were 86$\%$ congruent with morphology while the values were 89$\%$ for OC, 74$\%$ for PTP, and 72$\%$ for ABGD. [Biodiversity discovery; dark taxa; DNA barcodes; integrative taxonomy.].
© The Author(s) 2022. Published by Oxford University Press on behalf of the Society of Systematic Biologists.
Figures





References
-
- Ahrens D., Fujisawa T., Krammer H.-J., Eberle J., Fabrizi S., Vogler A.P.. 2016. Rarity and incomplete sampling in DNA-based species delimitation. Syst. Biol. 65:17. - PubMed
-
- Andersen A., Simcox D.J., Thomas J.A., Nash D.R.. 2014. Assessing reintroduction schemes by comparing genetic diversity of reintroduced and source populations: a case study of the globally threatened large blue butterfly (Maculinea arion). Biol. Conserv. 175:34–41.
-
- Bickel D. 2009. Why Hilara is not amusing: the problem of open-ended taxa and the limits of taxonomic knowledge. Diptera diversity: status, challenges, and tools. Leiden, Netherlands: E. J. Brill. p. 279–301.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

