Essential role of Cp190 in physical and regulatory boundary formation
- PMID: 35559678
- PMCID: PMC9106302
- DOI: 10.1126/sciadv.abl8834
Essential role of Cp190 in physical and regulatory boundary formation
Abstract
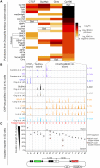
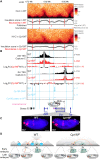
Boundaries in animal genomes delimit contact domains with enhanced internal contact frequencies and have debated functions in limiting regulatory cross-talk between domains and guiding enhancers to target promoters. Most mammalian boundaries form by stalling of chromosomal loop-extruding cohesin by CTCF, but most Drosophila boundaries form CTCF independently. However, how CTCF-independent boundaries form and function remains largely unexplored. Here, we assess genome folding and developmental gene expression in fly embryos lacking the ubiquitous boundary-associated factor Cp190. We find that sequence-specific DNA binding proteins such as CTCF and Su(Hw) directly interact with and recruit Cp190 to form most promoter-distal boundaries. Cp190 is essential for early development and prevents regulatory cross-talk between specific gene loci that pattern the embryo. Cp190 was, in contrast, dispensable for long-range enhancer-promoter communication at tested loci. Cp190 is thus currently the major player in fly boundary formation and function, revealing that diverse mechanisms evolved to partition genomes into independent regulatory domains.
Figures





References
-
- Rao S. S. P., Huang S.-C., Hilaire B. G. S., Engreitz J. M., Perez E. M., Kieffer-Kwon K.-R., Sanborn A. L., Johnstone S. E., Bascom G. D., Bochkov I. D., Huang X., Shamim M. S., Shin J., Turner D., Ye Z., Omer A. D., Robinson J. T., Schlick T., Bernstein B. E., Casellas R., Lander E. S., Aiden E. L., Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017). - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

