Detecting anti-SARS-CoV-2 antibodies in urine samples: A noninvasive and sensitive way to assay COVID-19 immune conversion
- PMID: 35559681
- PMCID: PMC9106288
- DOI: 10.1126/sciadv.abn7424
Detecting anti-SARS-CoV-2 antibodies in urine samples: A noninvasive and sensitive way to assay COVID-19 immune conversion
Abstract
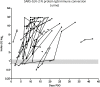
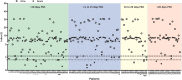
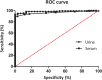
Serum-based ELISA (enzyme-linked immunosorbent assay) has been widely used to detect anti-severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) antibodies. However, to date, no study has investigated patient urine as a biological sample to detect SARS-CoV-2 virus-specific antibodies. An in-house urine-based ELISA was developed using recombinant SARS-CoV-2 nucleocapsid protein. The presence of SARS-CoV-2 antibodies in urine was established, with 94% sensitivity and 100% specificity for the detection of anti-SARS-CoV-2 antibodies with the urine-based ELISA and 88% sensitivity and 100% specificity with a paired serum-based ELISA. The urine-based ELISA that detects anti-SARS-CoV-2 antibodies is a noninvasive method with potential application as a facile COVID-19 immunodiagnostic platform, which can be used to report the extent of exposure at the population level and/or to assess the risk of infection at the individual level.
Figures

References
-
- World Health Organization, Coronavirus (COVID-19) dashboard (World Health Organization, 2021); https://COVID19.who.int/.
-
- World Health Organization, Antigen-detection in the diagnosis of SARS-CoV-2 infection (World Health Organization, 2021); www.who.int/publications/i/item/antigen-detection-in-the-diagnosis-of-sa....
-
- Yang Y., Yang M., Yuan J., Wang F., Wang Z., Li J., Zhang M., Xing L., Wei J., Peng L., Wong G., Zheng H., Wu W., Shen C., Liao M., Feng K., Li J., Yang Q., Zhao J., Liu L., Liu Y., Laboratory diagnosis and monitoring the viral shedding of SARS-CoV-2 infection. Innovation 1, 100061 (2020). - PMC - PubMed
-
- Weissleder R., Lee H., Ko J., Pittet M. J., COVID-19 diagnostics in context. Sci. Transl. Med. 12, eabc1931 (2020). - PubMed
-
- Amanat F., Stadlbauer D., Strohmeier S., Nguyen T. H. O., Chromikova V., McMahon M., Jiang K., Arunkumar G. A., Jurczyszak D., Polanco J., Bermudez-Gonzalez M., Kleiner G., Aydillo T., Miorin L., Fierer D. S., Lugo L. A., Kojic E. M., Stoever J., Liu S. T. H., Cunningham-Rundles C., Felgner P. L., Moran T., García-Sastre A., Caplivski D., Cheng A. C., Kedzierska K., Vapalahti O., Hepojoki J. M., Simon V., Krammer F., A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020). - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

