PTNet3D: A 3D High-Resolution Longitudinal Infant Brain MRI Synthesizer Based on Transformers
- PMID: 35560070
- PMCID: PMC9529847
- DOI: 10.1109/TMI.2022.3174827
PTNet3D: A 3D High-Resolution Longitudinal Infant Brain MRI Synthesizer Based on Transformers
Abstract
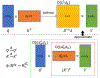
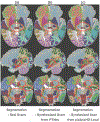
An increased interest in longitudinal neurodevelopment during the first few years after birth has emerged in recent years. Noninvasive magnetic resonance imaging (MRI) can provide crucial information about the development of brain structures in the early months of life. Despite the success of MRI collections and analysis for adults, it remains a challenge for researchers to collect high-quality multimodal MRIs from developing infant brains because of their irregular sleep pattern, limited attention, inability to follow instructions to stay still during scanning. In addition, there are limited analytic approaches available. These challenges often lead to a significant reduction of usable MRI scans and pose a problem for modeling neurodevelopmental trajectories. Researchers have explored solving this problem by synthesizing realistic MRIs to replace corrupted ones. Among synthesis methods, the convolutional neural network-based (CNN-based) generative adversarial networks (GANs) have demonstrated promising performance. In this study, we introduced a novel 3D MRI synthesis framework- pyramid transformer network (PTNet3D)- which relies on attention mechanisms through transformer and performer layers. We conducted extensive experiments on high-resolution Developing Human Connectome Project (dHCP) and longitudinal Baby Connectome Project (BCP) datasets. Compared with CNN-based GANs, PTNet3D consistently shows superior synthesis accuracy and superior generalization on two independent, large-scale infant brain MRI datasets. Notably, we demonstrate that PTNet3D synthesized more realistic scans than CNN-based models when the input is from multi-age subjects. Potential applications of PTNet3D include synthesizing corrupted or missing images. By replacing corrupted scans with synthesized ones, we observed significant improvement in infant whole brain segmentation.
Figures

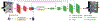







Similar articles
-
A 4D infant brain volumetric atlas based on the UNC/UMN baby connectome project (BCP) cohort.Neuroimage. 2022 Jun;253:119097. doi: 10.1016/j.neuroimage.2022.119097. Epub 2022 Mar 14. Neuroimage. 2022. PMID: 35301130 Free PMC article.
-
Transformer based multi-modal MRI fusion for prediction of post-menstrual age and neonatal brain development analysis.Med Image Anal. 2024 May;94:103140. doi: 10.1016/j.media.2024.103140. Epub 2024 Mar 7. Med Image Anal. 2024. PMID: 38461655
-
3D-MASNet: 3D mixed-scale asymmetric convolutional segmentation network for 6-month-old infant brain MR images.Hum Brain Mapp. 2023 Mar;44(4):1779-1792. doi: 10.1002/hbm.26174. Epub 2022 Dec 14. Hum Brain Mapp. 2023. PMID: 36515219 Free PMC article.
-
Resting-state functional MRI studies on infant brains: A decade of gap-filling efforts.Neuroimage. 2019 Jan 15;185:664-684. doi: 10.1016/j.neuroimage.2018.07.004. Epub 2018 Jul 7. Neuroimage. 2019. PMID: 29990581 Free PMC article. Review.
-
The UNC/UMN Baby Connectome Project (BCP): An overview of the study design and protocol development.Neuroimage. 2019 Jan 15;185:891-905. doi: 10.1016/j.neuroimage.2018.03.049. Epub 2018 Mar 22. Neuroimage. 2019. PMID: 29578031 Free PMC article. Review.
Cited by
-
One Model to Synthesize Them All: Multi-Contrast Multi-Scale Transformer for Missing Data Imputation.IEEE Trans Med Imaging. 2023 Sep;42(9):2577-2591. doi: 10.1109/TMI.2023.3261707. Epub 2023 Aug 31. IEEE Trans Med Imaging. 2023. PMID: 37030684 Free PMC article.
-
LungViT: Ensembling Cascade of Texture Sensitive Hierarchical Vision Transformers for Cross-Volume Chest CT Image-to-Image Translation.IEEE Trans Med Imaging. 2024 Jul;43(7):2448-2465. doi: 10.1109/TMI.2024.3367321. Epub 2024 Jul 1. IEEE Trans Med Imaging. 2024. PMID: 38373126 Free PMC article.
-
Unsupervised single-image super-resolution for infant brain MRI.Neuroimage. 2025 Aug 15;317:121293. doi: 10.1016/j.neuroimage.2025.121293. Epub 2025 Jun 21. Neuroimage. 2025. PMID: 40550405 Free PMC article.
-
SCANeXt: Enhancing 3D medical image segmentation with dual attention network and depth-wise convolution.Heliyon. 2024 Feb 28;10(5):e26775. doi: 10.1016/j.heliyon.2024.e26775. eCollection 2024 Mar 15. Heliyon. 2024. PMID: 38439873 Free PMC article.
-
Synthesizing pseudo-T2w images to recapture missing data in neonatal neuroimaging with applications in rs-fMRI.Neuroimage. 2022 Jun;253:119091. doi: 10.1016/j.neuroimage.2022.119091. Epub 2022 Mar 11. Neuroimage. 2022. PMID: 35288282 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

