The Quasimesenchymal Pancreatic Ductal Epithelial Cell Line PANC-1-A Useful Model to Study Clonal Heterogeneity and EMT Subtype Shifting
- PMID: 35565186
- PMCID: PMC9101310
- DOI: 10.3390/cancers14092057
The Quasimesenchymal Pancreatic Ductal Epithelial Cell Line PANC-1-A Useful Model to Study Clonal Heterogeneity and EMT Subtype Shifting
Abstract
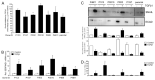
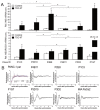
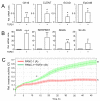
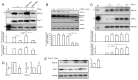
Intratumoral heterogeneity (ITH) is an intrinsic feature of malignant tumors that eventually allows a subfraction of resistant cancer cells to clonally evolve and cause therapy failure or relapse. ITH, cellular plasticity and tumor progression are driven by epithelial-mesenchymal transition (EMT) and the reverse process, MET. During these developmental programs, epithelial (E) cells are successively converted to invasive mesenchymal (M) cells, or back to E cells, by passing through a series of intermediate E/M states, a phenomenon termed E-M plasticity (EMP). The induction of MET has clinical potential as it can block the initial EMT stages that favor tumor cell dissemination, while its inhibition can curb metastatic outgrowth at distant sites. In pancreatic ductal adenocarcinoma (PDAC), cellular models with which to study EMP or MET induction are scarce. Here, we have generated single cell-derived clonal cultures of the quasimesenchymal PDAC-derived cell line, PANC-1, and found that these differ strongly with respect to cell morphology and EMT marker expression, allowing for their tentative classification as E, E/M or M. Interestingly, the different EMT phenotypes were found to segregate with differences in tumorigenic potential in vitro, as measured by colony forming and invasive activities, and in circadian clock function. Moreover, the individual clones the phenotypes of which remained stable upon prolonged culture also responded differently to treatment with transforming growth factor (TGF)β1 in regard to regulation of growth and individual TGFβ target genes, and to culture conditions that favour ductal-to-endocrine transdifferentiation as a more direct measure for cellular plasticity. Of note, stimulation with TGFβ1 induced a shift in parental PANC-1 cultures towards a more extreme M and invasive phenotype, while exposing the cells to a combination of the proinflammatory cytokines IFNγ, IL1β and TNFα (IIT) elicited a shift towards a more E and less invasive phenotype resembling a MET-like process. Finally, we show that the actions of TGFβ1 and IIT both converge on regulating the ratio of the small GTPase RAC1 and its splice isoform, RAC1b. Our data provide strong evidence for dynamic EMT-MET transitions and qualify this cell line as a useful model with which to study EMP.
Keywords: EMT; MET; PANC-1; epithelial–mesenchymal plasticity; intratumoral heterogeneity; pancreatic ductal adenocarcinoma; phenotype; proinflammatory cytokines; transdifferentiation; transforming growth factor β.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Characterization of Epithelial-Mesenchymal and Neuroendocrine Differentiation States in Pancreatic and Small Cell Ovarian Tumor Cells and Their Modulation by TGF-β1 and BMP-7.Cells. 2024 Dec 5;13(23):2010. doi: 10.3390/cells13232010. Cells. 2024. PMID: 39682758 Free PMC article.
-
Activation of a Ductal-to-Endocrine Transdifferentiation Transcriptional Program in the Pancreatic Cancer Cell Line PANC-1 Is Controlled by RAC1 and RAC1b through Antagonistic Regulation of Stemness Factors.Cancers (Basel). 2021 Nov 4;13(21):5541. doi: 10.3390/cancers13215541. Cancers (Basel). 2021. PMID: 34771704 Free PMC article.
-
Rac1b negatively regulates TGF-β1-induced cell motility in pancreatic ductal epithelial cells by suppressing Smad signalling.Oncotarget. 2014 Jan 15;5(1):277-90. doi: 10.18632/oncotarget.1696. Oncotarget. 2014. PMID: 24378395 Free PMC article.
-
Epithelial to Mesenchymal Transition: Key Regulator of Pancreatic Ductal Adenocarcinoma Progression and Chemoresistance.Cancers (Basel). 2021 Nov 4;13(21):5532. doi: 10.3390/cancers13215532. Cancers (Basel). 2021. PMID: 34771695 Free PMC article. Review.
-
The Role of SMAD4 Inactivation in Epithelial-Mesenchymal Plasticity of Pancreatic Ductal Adenocarcinoma: The Missing Link?Cancers (Basel). 2022 Feb 15;14(4):973. doi: 10.3390/cancers14040973. Cancers (Basel). 2022. PMID: 35205719 Free PMC article. Review.
Cited by
-
Elucidation of the Role of SMAD4 in Epithelial-Mesenchymal Plasticity: Does It Help to Better Understand the Consequences of DPC4 Inactivation in the Malignant Progression of Pancreatic Ductal Adenocarcinoma?Cancers (Basel). 2023 Jan 18;15(3):581. doi: 10.3390/cancers15030581. Cancers (Basel). 2023. PMID: 36765537 Free PMC article.
-
PANC-1 Cell Line as an Experimental Model for Characterizing PIVKA-II Production, Distribution, and Molecular Mechanisms Leading to Protein Release in PDAC.Int J Mol Sci. 2024 Mar 20;25(6):3498. doi: 10.3390/ijms25063498. Int J Mol Sci. 2024. PMID: 38542466 Free PMC article.
-
Multifaceted Effects of Kinase Inhibitors on Pancreatic Cancer Cells Reveals Pivotal Entities with Therapeutic Implications.Biomedicines. 2023 Jun 15;11(6):1716. doi: 10.3390/biomedicines11061716. Biomedicines. 2023. PMID: 37371811 Free PMC article.
-
The Oncoprotein Mucin 1 in Pancreatic Cancer Onset and Progression: Potential Clinical Implications.Biomolecules. 2025 Feb 13;15(2):275. doi: 10.3390/biom15020275. Biomolecules. 2025. PMID: 40001578 Free PMC article. Review.
-
Emerging roles of endoplasmic reticulum stress in the cellular plasticity of cancer cells.Front Oncol. 2023 Feb 20;13:1110881. doi: 10.3389/fonc.2023.1110881. eCollection 2023. Front Oncol. 2023. PMID: 36890838 Free PMC article. Review.
References
-
- Katsuno Y., Meyer D.S., Zhang Z., Shokat K.M., Akhurst R.J., Miyazono K., Dernyck R. Chronic TGF-β exposure drives stabilized EMT, tumor stemness, and cancer drug resistance with vulnerability to bitopic mTOR inhibition. Sci. Signal. 2019;12:eaau8544. doi: 10.1126/scisignal.aau8544. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

