Machine-Learning-Based Clinical Biomarker Using Cell-Free DNA for Hepatocellular Carcinoma (HCC)
- PMID: 35565192
- PMCID: PMC9103537
- DOI: 10.3390/cancers14092061
Machine-Learning-Based Clinical Biomarker Using Cell-Free DNA for Hepatocellular Carcinoma (HCC)
Abstract
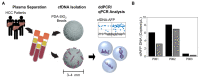
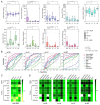
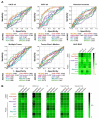
(1) Background: Hepatocellular carcinoma (HCC) is one of the leading causes of cancer-related death worldwide. Although various serum enzymes have been utilized for the diagnosis and prognosis of HCC, the currently available biomarkers lack the sensitivity needed to detect HCC at early stages and accurately predict treatment responses. (2) Methods: We utilized our highly sensitive cell-free DNA (cfDNA) detection system, in combination with a machine learning algorithm, to provide a platform for improved diagnosis and prognosis of HCC. (3) Results: cfDNA, specifically alpha-fetoprotein (AFP) expression in captured cfDNA, demonstrated the highest accuracy for diagnosing malignancies among the serum/plasma biomarkers used in this study, including AFP, aspartate aminotransferase, alanine aminotransferase, albumin, alkaline phosphatase, and bilirubin. The diagnostic/prognostic capability of cfDNA was further improved by establishing a cfDNA score (cfDHCC), which integrated the total plasma cfDNA levels and cfAFP-DNA expression into a single score using machine learning algorithms. (4) Conclusion: The cfDHCC score demonstrated significantly improved accuracy in determining the pathological features of HCC and predicting patients' survival outcomes compared to the other biomarkers. The results presented herein reveal that our cfDNA capture/analysis platform is a promising approach to effectively utilize cfDNA as a biomarker for the diagnosis and prognosis of HCC.
Keywords: cell-free DNA (cfDNA); circulating tumor DNA (ctDNA); hepatocellular carcinoma (HCC); liquid biopsy; principal component analysis (PCA).
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Utility of Liquid Biopsy Analysis in Detection of Hepatocellular Carcinoma, Determination of Prognosis, and Disease Monitoring: A Systematic Review.Clin Gastroenterol Hepatol. 2020 Dec;18(13):2879-2902.e9. doi: 10.1016/j.cgh.2020.04.019. Epub 2020 Apr 11. Clin Gastroenterol Hepatol. 2020. PMID: 32289533 Free PMC article.
-
Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy.Cancers (Basel). 2022 Jul 11;14(14):3367. doi: 10.3390/cancers14143367. Cancers (Basel). 2022. PMID: 35884434 Free PMC article.
-
Diagnostic value of circulating cell-free DNA levels for hepatocellular carcinoma.Int J Infect Dis. 2018 Feb;67:92-97. doi: 10.1016/j.ijid.2017.12.002. Epub 2017 Dec 8. Int J Infect Dis. 2018. PMID: 29229500 Clinical Trial.
-
Liquid biopsy using cell-free DNA in the early diagnosis of hepatocellular carcinoma.Invest New Drugs. 2023 Jun;41(3):532-538. doi: 10.1007/s10637-023-01363-6. Epub 2023 Apr 26. Invest New Drugs. 2023. PMID: 37099161 Review.
-
Detection of a novel panel of somatic mutations in plasma cell-free DNA and its diagnostic value in hepatocellular carcinoma.Cancer Manag Res. 2019 Jun 28;11:5745-5756. doi: 10.2147/CMAR.S197455. eCollection 2019. Cancer Manag Res. 2019. PMID: 31303788 Free PMC article.
Cited by
-
Current and Future Perspectives of Cell-Free DNA in Liquid Biopsy.Curr Issues Mol Biol. 2022 Jun 10;44(6):2695-2709. doi: 10.3390/cimb44060184. Curr Issues Mol Biol. 2022. PMID: 35735625 Free PMC article. Review.
-
Dual Biomarker Strategies for Liquid Biopsy: Integrating Circulating Tumor Cells and Circulating Tumor DNA for Enhanced Tumor Monitoring.Biosensors (Basel). 2025 Jan 28;15(2):74. doi: 10.3390/bios15020074. Biosensors (Basel). 2025. PMID: 39996976 Free PMC article. Review.
-
Biomarkers for diagnosis and therapeutic options in hepatocellular carcinoma.Mol Cancer. 2024 Sep 6;23(1):189. doi: 10.1186/s12943-024-02101-z. Mol Cancer. 2024. PMID: 39242496 Free PMC article. Review.
-
Advances in novel strategies for isolation, characterization, and analysis of CTCs and ctDNA.Ther Adv Med Oncol. 2023 Sep 7;15:17588359231192401. doi: 10.1177/17588359231192401. eCollection 2023. Ther Adv Med Oncol. 2023. PMID: 37692363 Free PMC article. Review.
-
Programmed Death 1 and Cytotoxic T-Lymphocyte-Associated Protein 4 Gene Expression in Peripheral Blood Mononuclear Cells Can Serve as Prognostic Biomarkers for Hepatocellular Carcinoma.Cancers (Basel). 2024 Apr 13;16(8):1493. doi: 10.3390/cancers16081493. Cancers (Basel). 2024. PMID: 38672574 Free PMC article.
References
-
- Akinyemiju T., Abera S., Ahmed M., Alam N., Alemayohu M.A., Allen C., Al-Raddadi R., Alvis-Guzman N., Amoako Y., Artaman A., et al. The Burden of Primary Liver Cancer and Underlying Etiologies from 1990 to 2015 at the Global, Regional, and National Level: Results from the Global Burden of Disease Study 2015. JAMA Oncol. 2017;3:1683–1691. - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

